⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_00694
- pan locus tag?: SAUPAN002570000
- symbol: SAOUHSC_00694
- pan gene symbol?: mgrA
- synonym:
- product: hypothetical protein
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_00694
- symbol: SAOUHSC_00694
- product: hypothetical protein
- replicon: chromosome
- strand: -
- coordinates: 679331..679774
- length: 444
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3920998 NCBI
- RefSeq: YP_499253 NCBI
- BioCyc: G1I0R-648 BioCyc
- MicrobesOnline: 1289163 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421ATGTCTGATCAACATAATTTAAAAGAACAGCTATGCTTTAGTTTGTACAATGCTCAAAGA
CAAGTTAATCGCTACTACTCTAACAAAGTTTTTAAGAAGTACAATCTAACATACCCACAA
TTTCTTGTCTTAACAATTTTATGGGATGAATCTCCTGTAAACGTCAAGAAAGTCGTAACT
GAATTAGCACTCGATACTGGTACAGTATCACCATTATTAAAACGAATGGAACAAGTAGAC
TTAATTAAGCGTGAACGTTCCGAAGTCGATCAACGTGAAGTATTTATTCACTTGACTGAC
AAAAGTGAAACTATTAGACCAGAATTAAGTAATGCATCTGACAAAGTCGCTTCAGCTTCT
TCTTTATCGCAAGATGAAGTTAAAGAACTTAATCGCTTATTAGGTAAAGTCATTCATGCA
TTTGATGAAACAAAGGAAAAATAA60
120
180
240
300
360
420
444
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_00694
- symbol: SAOUHSC_00694
- description: hypothetical protein
- length: 147
- theoretical pI: 7.59928
- theoretical MW: 17089.3
- GRAVY: -0.564626
⊟Function[edit | edit source]
- TIGRFAM: mobile rSAM pair MarR family regulator (TIGR04472; HMM-score: 54.1)homoprotocatechuate degradation operon regulator, HpaR (TIGR02337; HMM-score: 51.6)Regulatory functions DNA interactions staphylococcal accessory regulator family (TIGR01889; HMM-score: 47.2)
- TheSEED :
- Transcriptional regulator MgrA (Regulator of autolytic activity)
- ⊞PFAM: HTH (CL0123) MarR; MarR family (PF01047; HMM-score: 53.9)and 6 more
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MSDQHNLKEQLCFSLYNAQRQVNRYYSNKVFKKYNLTYPQFLVLTILWDESPVNVKKVVTELALDTGTVSPLLKRMEQVDLIKRERSEVDQREVFIHLTDKSETIRPELSNASDKVASASSLSQDEVKELNRLLGKVIHAFDETKEK
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas [1] [2]
- protein localization: data available for COL
- quantitative data / protein copy number per cell: data available for COL
- ⊟interaction partners:
SAOUHSC_01757 (rplU) 50S ribosomal protein L21 [3] (data from MRSA252) SAOUHSC_00196 hypothetical protein [3] (data from MRSA252) SAOUHSC_01416 dihydrolipoamide succinyltransferase [3] (data from MRSA252)
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- predicted SigA promoter [4] : S264 < SAOUHSC_00694 < S266
⊟Regulation[edit | edit source]
- ⊞regulator: RNAIII/S871 regulon
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [4]
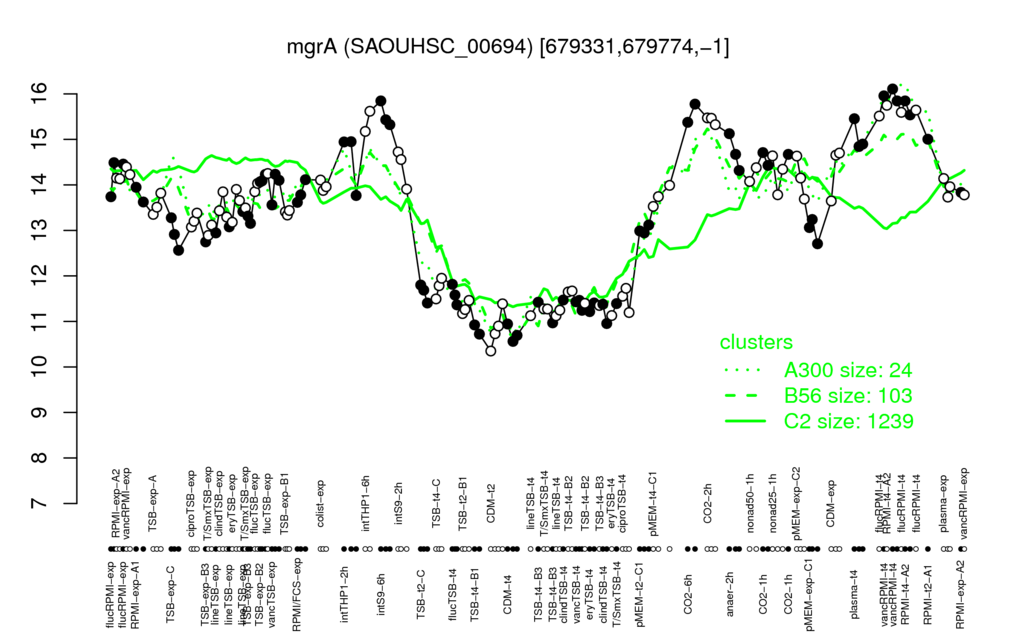 ⊟Multi-gene expression profiles
⊟Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊞Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊞Other Information[edit | edit source]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Maren Depke, Stephan Michalik, Alexander Rabe, Kristin Surmann, Lars Brinkmann, Nico Jehmlich, Jörg Bernhardt, Michael Hecker, Bernd Wollscheid, Zhi Sun, Robert L Moritz, Uwe Völker, Frank Schmidt
A peptide resource for the analysis of Staphylococcus aureus in host-pathogen interaction studies.
Proteomics: 2015, 15(21);3648-61
[PubMed:26224020] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Maren Depke, Annette Murr, Manuela Gesell Salazar, Ulrike Kusebauch, Zhi Sun, Tanja C Meyer, Kristin Surmann, Henrike Pförtner, Petra Hildebrandt, Stefan Weiss, Laura Marcela Palma Medina, Melanie Gutjahr, Elke Hammer, Dörte Becher, Thomas Pribyl, Sven Hammerschmidt, Eric W Deutsch, Samuel L Bader, Michael Hecker, Robert L Moritz, Ulrike Mäder, Uwe Völker, Frank Schmidt
A global Staphylococcus aureus proteome resource applied to the in vivo characterization of host-pathogen interactions.
Sci Rep: 2017, 7(1);9718
[PubMed:28887440] [WorldCat.org] [DOI] (I e) - ↑ Jump up to: 3.0 3.1 3.2 Artem Cherkasov, Michael Hsing, Roya Zoraghi, Leonard J Foster, Raymond H See, Nikolay Stoynov, Jihong Jiang, Sukhbir Kaur, Tian Lian, Linda Jackson, Huansheng Gong, Rick Swayze, Emily Amandoron, Farhad Hormozdiari, Phuong Dao, Cenk Sahinalp, Osvaldo Santos-Filho, Peter Axerio-Cilies, Kendall Byler, William R McMaster, Robert C Brunham, B Brett Finlay, Neil E Reiner
Mapping the protein interaction network in methicillin-resistant Staphylococcus aureus.
J Proteome Res: 2011, 10(3);1139-50
[PubMed:21166474] [WorldCat.org] [DOI] (I p) - ↑ Jump up to: 4.0 4.1 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e) - ↑ Delphine Bronesky, Zongfu Wu, Stefano Marzi, Philippe Walter, Thomas Geissmann, Karen Moreau, François Vandenesch, Isabelle Caldelari, Pascale Romby
Staphylococcus aureus RNAIII and Its Regulon Link Quorum Sensing, Stress Responses, Metabolic Adaptation, and Regulation of Virulence Gene Expression.
Annu Rev Microbiol: 2016, 70;299-316
[PubMed:27482744] [WorldCat.org] [DOI] (I p)
⊟Relevant publications[edit | edit source]
Que Chi Truong-Bolduc, Xiamei Zhang, David C Hooper
Characterization of NorR protein, a multifunctional regulator of norA expression in Staphylococcus aureus.
J Bacteriol: 2003, 185(10);3127-38
[PubMed:12730173] [WorldCat.org] [DOI] (P p)S S Ingavale, W Van Wamel, A L Cheung
Characterization of RAT, an autolysis regulator in Staphylococcus aureus.
Mol Microbiol: 2003, 48(6);1451-66
[PubMed:12791130] [WorldCat.org] [DOI] (P p)Adhar C Manna, Susham S Ingavale, MaryBeth Maloney, Willem van Wamel, Ambrose L Cheung
Identification of sarV (SA2062), a new transcriptional regulator, is repressed by SarA and MgrA (SA0641) and involved in the regulation of autolysis in Staphylococcus aureus.
J Bacteriol: 2004, 186(16);5267-80
[PubMed:15292128] [WorldCat.org] [DOI] (P p)Susham Ingavale, Willem van Wamel, Thanh T Luong, Chia Y Lee, Ambrose L Cheung
Rat/MgrA, a regulator of autolysis, is a regulator of virulence genes in Staphylococcus aureus.
Infect Immun: 2005, 73(3);1423-31
[PubMed:15731040] [WorldCat.org] [DOI] (P p)Q C Truong-Bolduc, P M Dunman, J Strahilevitz, S J Projan, D C Hooper
MgrA is a multiple regulator of two new efflux pumps in Staphylococcus aureus.
J Bacteriol: 2005, 187(7);2395-405
[PubMed:15774883] [WorldCat.org] [DOI] (P p)
