Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_00509
- pan locus tag?: SAUPAN002294000
- symbol: gltX
- pan gene symbol?: gltX
- synonym:
- product: glutamyl-tRNA synthetase
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
⊟Accession numbers[edit | edit source]
- Gene ID: 3920421 NCBI
- RefSeq: YP_499082 NCBI
- BioCyc: G1I0R-480 BioCyc
- MicrobesOnline: 1288992 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201
1261
1321
1381
1441ATGAGCGATCGTATAAGAGTAAGATATGCACCAAGTCCAACTGGGTATCTTCATATTGGT
AATGCAAGAACAGCATTATTCAATTACTTGTATGCTAAACATTACAACGGAGATTTTGTG
ATTCGAATTGAAGATACTGATAAAAAACGTAATTTAGAAGATGGAGAAACATCACAATTT
GATAATCTTAAATGGTTAGGATTAGATTGGGATGAGTCTGTAGATAAAGACAATGGCTAC
GGACCATATCGTCAATCTGAACGTCAACATATCTACCAACCATTAATAGATCAGTTACTA
GCAGAAGATAAAGCATATAAATGCTATATGACAGAAGAAGAATTAGAAGCTGAACGTGAA
GCGCAAATCGCTCGTGGTGAAATGCCTCGCTATGGTGGTCAACATGCGCATTTGACTGAA
GAACAACGTCAACAATTTGAAGCAGAAGGACGCCAACCATCAATTCGTTTCCGAGTACCT
CAAAACCAAACGTATTCATTTGATGATATGGTAAAAGGAAATATTTCATTTGATTCAAAT
GGTATTGGTGACTGGGTTATCGTAAAAAAAGATGGCATTCCAACGTACAATTTTGCAGTA
GCTATAGATGATCATTACATGCAAATTTCAGATGTAATTCGTGGTGATGATCATATTTCA
AACACGCCTAAACAAATTATGATTTATGAAGCATTTGGCTGGGAGCCACCTCGTTTTGGT
CATATGTCATTAATTGTTAATGAAGAACGTAAAAAGTTAAGTAAACGTGATGGGCAAATT
TTACAATTTATTGAGCAATATCGTGACTTAGGTTATTTACCTGAAGCGTTATTTAATTTT
ATTGCGTTATTAGGTTGGTCTCCTGAAGGTGAAGAAGAAATCTTTTCTAAAGAAGAATTT
ATCAAAATCTTTGATGAAAAGCGTTTGTCAAAATCACCAGCATTTTTCGATAAGCAAAAA
TTAGCATGGGTTAATAACCAATATATGAAACAAAAAGATACTGAAACAGTATTCCAATTA
GCATTACCTCATTTAATTAAAGCAAATTTGATTCCTGAGGTGCCGTCAGAAGAGGATTTA
TCTTGGGGACGCAAATTAATTGCGCTTTATCAAAAAGAAATGAGTTATGCCGGTGAAATT
GTACCTTTATCAGAAATGTTCTTTAAAGAAATGCCAGCTCTTGGTGAAGAAGAACAACAA
GTGATTAATGGAGAGCAAGTACCAGAGTTAATGACGCACTTATTCAGTAAATTAGAAGCA
CTTGAACCATTTGAAGCGGCTGAAATTAAAAAGACAATTAAAGAAGTTCAAAAAGAAACA
GGAATAAAAGGCAAGCAATTATTTATGCCTATTCGTGTTGCTGTAACAGGCCAAATGCAT
GGTCCTGAATTACCAAATACAATTGAAGTACTTGGTAAAGAAAAAGTGCTAAACCGTTTA
AAACAATATAAGTAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1260
1320
1380
1440
1455
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_00509
- symbol: GltX
- description: glutamyl-tRNA synthetase
- length: 484
- theoretical pI: 4.97567
- theoretical MW: 56288.4
- GRAVY: -0.639669
⊟Function[edit | edit source]
- reaction: EC 6.1.1.17? ExPASyGlutamate--tRNA ligase ATP + L-glutamate + tRNA(Glu) = AMP + diphosphate + L-glutamyl-tRNA(Glu)
- TIGRFAM: Protein synthesis tRNA aminoacylation glutamate--tRNA ligase (TIGR00464; EC 6.1.1.17; HMM-score: 624.4)and 4 moreProtein synthesis tRNA and rRNA base modification glutamyl-queuosine tRNA(Asp) synthetase (TIGR03838; EC 6.1.1.-; HMM-score: 219.3)Protein synthesis tRNA aminoacylation glutamate--tRNA ligase (TIGR00463; EC 6.1.1.17; HMM-score: 127.7)Protein synthesis tRNA aminoacylation glutamine--tRNA ligase (TIGR00440; EC 6.1.1.18; HMM-score: 52.3)Protein synthesis tRNA aminoacylation lysine--tRNA ligase (TIGR00467; EC 6.1.1.6; HMM-score: 28.1)
- TheSEED :
- tRNA glutamyl-Q(34)-Gln synthetase (EC 6.1.1.24)
- tRNA glutamyl-Q(34)-Glu synthetase (EC 6.1.1.17)
Protein Metabolism Protein biosynthesis tRNA aminoacylation, Glu and Gln Glutamyl-tRNA synthetase (EC 6.1.1.17)and 3 moreProtein Metabolism Protein biosynthesis tRNA aminoacylation, Glu and Gln Glutamyl-tRNA(Gln) synthetase (EC 6.1.1.24) - PFAM: HUP (CL0039) tRNA-synt_1c; tRNA synthetases class I (E and Q), catalytic domain (PF00749; HMM-score: 412.1)and 4 moreno clan defined Anticodon_2; Anticodon binding domain (PF19269; HMM-score: 105.2)HUP (CL0039) tRNA-synt_1f; tRNA synthetases class I (K) (PF01921; HMM-score: 18.1)HTH (CL0123) Mrr_N; Mrr N-terminal domain (PF14338; HMM-score: 13.3)no clan defined VMAP-M5; vWA-MoxR associated protein middle region (VMAP-M) 5 (PF19967; HMM-score: 12.2)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 10
- Cytoplasmic Membrane Score: 0
- Cellwall Score: 0
- Extracellular Score: 0
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.9996
- Cytoplasmic Membrane Score: 0
- Cell wall & surface Score: 0
- Extracellular Score: 0.0003
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: -1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.003965
- TAT(Tat/SPI): 0.000323
- LIPO(Sec/SPII): 0.000767
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MSDRIRVRYAPSPTGYLHIGNARTALFNYLYAKHYNGDFVIRIEDTDKKRNLEDGETSQFDNLKWLGLDWDESVDKDNGYGPYRQSERQHIYQPLIDQLLAEDKAYKCYMTEEELEAEREAQIARGEMPRYGGQHAHLTEEQRQQFEAEGRQPSIRFRVPQNQTYSFDDMVKGNISFDSNGIGDWVIVKKDGIPTYNFAVAIDDHYMQISDVIRGDDHISNTPKQIMIYEAFGWEPPRFGHMSLIVNEERKKLSKRDGQILQFIEQYRDLGYLPEALFNFIALLGWSPEGEEEIFSKEEFIKIFDEKRLSKSPAFFDKQKLAWVNNQYMKQKDTETVFQLALPHLIKANLIPEVPSEEDLSWGRKLIALYQKEMSYAGEIVPLSEMFFKEMPALGEEEQQVINGEQVPELMTHLFSKLEALEPFEAAEIKKTIKEVQKETGIKGKQLFMPIRVAVTGQMHGPELPNTIEVLGKEKVLNRLKQYK
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas [2] [3]
- protein localization: data available for COL
- quantitative data / protein copy number per cell: data available for COL
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- predicted SigA promoter [4] : SAOUHSC_00502 > S169 > SAOUHSC_00503 > SAOUHSC_00504 > SAOUHSC_00505 > S170 > S171 > S172 > SAOUHSC_00507 > SAOUHSC_00508 > S173 > S174 > gltX > S175 > S176 > SAOUHSC_00510predicted SigA promoter [4] : S171 > S172 > SAOUHSC_00507 > SAOUHSC_00508 > S173 > S174 > gltX > S175 > S176 > SAOUHSC_00510 > cysS > SAOUHSC_00512 > SAOUHSC_00513 > SAOUHSC_00514 > S177 > SAOUHSC_00515 > S178 > S179 > secE > SAOUHSC_00517 > S180 > rplK > S181 > rplA > S182predicted SigA promoter [4] : S174 > gltX > S175 > S176 > SAOUHSC_00510 > cysS > SAOUHSC_00512 > SAOUHSC_00513 > SAOUHSC_00514 > S177 > SAOUHSC_00515 > S178 > S179 > secE > SAOUHSC_00517 > S180 > rplK > S181 > rplA > S182 > S183 > rplJ > rplL
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [4]
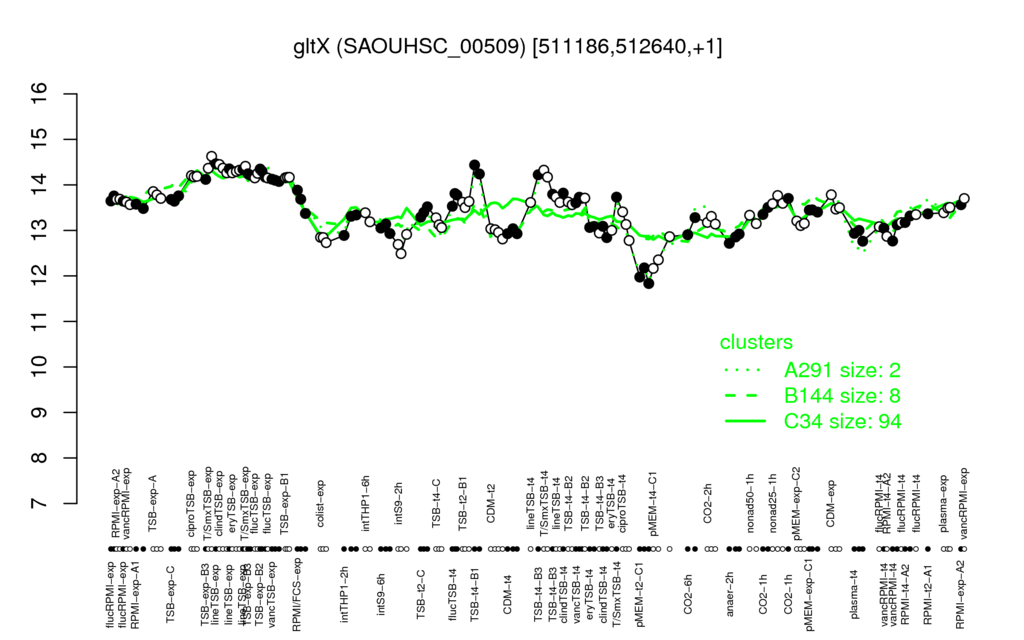 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Roy R Chaudhuri, Andrew G Allen, Paul J Owen, Gil Shalom, Karl Stone, Marcus Harrison, Timothy A Burgis, Michael Lockyer, Jorge Garcia-Lara, Simon J Foster, Stephen J Pleasance, Sarah E Peters, Duncan J Maskell, Ian G Charles
Comprehensive identification of essential Staphylococcus aureus genes using Transposon-Mediated Differential Hybridisation (TMDH).
BMC Genomics: 2009, 10;291
[PubMed:19570206] [WorldCat.org] [DOI] (I e) - ↑ Maren Depke, Stephan Michalik, Alexander Rabe, Kristin Surmann, Lars Brinkmann, Nico Jehmlich, Jörg Bernhardt, Michael Hecker, Bernd Wollscheid, Zhi Sun, Robert L Moritz, Uwe Völker, Frank Schmidt
A peptide resource for the analysis of Staphylococcus aureus in host-pathogen interaction studies.
Proteomics: 2015, 15(21);3648-61
[PubMed:26224020] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Maren Depke, Annette Murr, Manuela Gesell Salazar, Ulrike Kusebauch, Zhi Sun, Tanja C Meyer, Kristin Surmann, Henrike Pförtner, Petra Hildebrandt, Stefan Weiss, Laura Marcela Palma Medina, Melanie Gutjahr, Elke Hammer, Dörte Becher, Thomas Pribyl, Sven Hammerschmidt, Eric W Deutsch, Samuel L Bader, Michael Hecker, Robert L Moritz, Ulrike Mäder, Uwe Völker, Frank Schmidt
A global Staphylococcus aureus proteome resource applied to the in vivo characterization of host-pathogen interactions.
Sci Rep: 2017, 7(1);9718
[PubMed:28887440] [WorldCat.org] [DOI] (I e) - ↑ 4.0 4.1 4.2 4.3 4.4 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
