Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_01170
- pan locus tag?: SAUPAN003483000
- symbol: carB
- pan gene symbol?: carB
- synonym:
- product: carbamoyl phosphate synthase large subunit
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_01170
- symbol: carB
- product: carbamoyl phosphate synthase large subunit
- replicon: chromosome
- strand: +
- coordinates: 1118950..1122123
- length: 3174
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3920921 NCBI
- RefSeq: YP_499709 NCBI
- BioCyc: G1I0R-1096 BioCyc
- MicrobesOnline: 1289623 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201
1261
1321
1381
1441
1501
1561
1621
1681
1741
1801
1861
1921
1981
2041
2101
2161
2221
2281
2341
2401
2461
2521
2581
2641
2701
2761
2821
2881
2941
3001
3061
3121ATGCCTAAACGTAATGATATCAAAACAATTTTAGTAATAGGGTCTGGGCCAATTATCATA
GGTCAAGCAGCTGAATTTGATTATGCTGGAACACAAGCATGTCTAGCTTTAAAAGAAGAG
GGATATCGAGTTATTCTTGTAAATTCAAATCCAGCGACAATCATGACTGATAAGGAAATT
GCGGATAAAGTATATATCGAACCGTTAACTCATGATTTTATAGCGCGAATTATACGTAAA
GAGCAACCTGACGCTTTACTTCCAACTTTAGGTGGTCAAACAGGTTTAAACATGGCGATT
CAACTACACGAAAGTGGTGTGCTTCAAGATAATAACGTCCAATTATTAGGAACTGAGCTA
ACATCAATTCAACAAGCAGAAGACCGTGAAATGTTTAGAACATTAATGAATGATTTAAAC
GTTCCTGTACCAGAGAGTGACATTGTAAATACAGTAGAGCAAGCCTTTAAATTCAAAGAG
CAAGTGGGATACCCGCTAATTGTTAGACCGGCATTTACGATGGGTGGTACCGGAGGCGGT
ATTTGTCATAATGATGAAGAATTACATGAAATCGTCTCAAATGGTCTTCATTATAGTCCA
GCAACGCAATGTTTATTAGAAAAATCTATCGCAGGTTTTAAAGAAATCGAATACGAAGTA
ATGCGTGATAAAAACGATAATGCCATCGTTGTATGTAACATGGAAAATATTGATCCAGTT
GGTATTCATACAGGCGATTCAATTGTTGTGGCTCCTAGTCAAACATTATCAGATGTTGAG
TATCAAATGTTACGTGATGTTTCATTAAAAGTTATTCGAGCTTTAGGTATCGAAGGTGGT
TGTAATGTTCAATTAGCATTAGATCCCCATTCATTCGATTATTATATTATAGAAGTAAAT
CCGCGTGTATCACGTTCATCAGCGTTAGCTTCAAAAGCAACAGGATATCCTATTGCAAAA
TTAGCTGCTAAAATCGCGGTTGGTCTAACATTAGATGAAATGTTAAATCCAATTACAGGA
ACATCTTATGCAGCGTTTGAACCAACTTTAGACTATGTGATTTCAAAAATACCAAGATTT
CCTTTTGATAAATTTGAAAAAGGAGAACGAGAGCTTGGCACACAAATGAAAGCAACAGGT
GAAGTTATGGCCATTGGTCGAACTTACGAAGAATCATTGTTAAAAGCAATTCGATCACTT
GAGTATGGTGTGCATCACTTAGGATTACCAAATGGTGAAAGCTTCGATCTTGATTATATT
AAAGAACGTATTTCACACCAAGATGATGAACGATTATTTTTCATCGGCGAAGCAATTAGA
AGAGGCACAACATTAGAAGAAATTCATAATATGACTCAGATTGATTACTTCTTCTTACAC
AAGTTCCAAAACATTATTGATATTGAGCATCAATTAAAAGAGCATCAAGGTGATTTAGAA
TATCTTAAATATGCAAAAGATTATGGATTTAGTGATAAAACAATAGCGCATCGCTTTAAT
ATGACGGAAGAAGAAGTATATCAATTGCGTATGGAAAATGATATTAAACCTGTTTACAAG
ATGGTTGATACTTGCGCAGCTGAATTTGAATCTTCAACACCATATTATTATGGTACATAC
GAAACTGAAAATGAATCCATAGTTACTGACAAAGAAAAAATCTTAGTATTAGGCTCTGGA
CCAATTCGAATCGGCCAAGGTGTAGAATTTGACTATGCGACAGTTCACGCCGTTTGGGCA
ATTCAAAAAGCAGGGTACGAAGCGATAATTGTGAATAACAATCCAGAAACAGTTTCAACA
GACTTCTCAATTTCTGACAAATTATACTTTGAACCTTTAACTGAAGAAGATGTGATGAAT
ATCATTAATTTAGAAAAACCTAAAGGTGTCGTTGTACAATTTGGAGGACAAACAGCGATT
AATTTAGCAGACAAATTGGCTAAACATGGTGTTAAAATACTTGGTACTTCACTAGAAAAT
CTAAATCGTGCTGAAGATAGAAAAGAATTTGAAGCACTATTAAGAAAAATTAACGTGCCA
CAGCCACAAGGGAAAACAGCTACATCACCTGAGGAAGCATTAGCGAATGCTGCAGAAATC
GGATATCCGGTTGTAGTAAGACCTTCTTATGTATTAGGTGGTCGCGCAATGGAAATTGTA
GACAATGACAAAGAGTTAGAAAACTATATGACCCAGGCTGTAAAAGCGAGTCCGGAACAT
CCGGTACTAGTCGATAGATATTTAACTGGTAAAGAAATTGAAGTTGATGCGATTTGTGAT
GGAGAAACGGTCATTATTCCAGGAATCATGGAACATATTGAACGTGCTGGTGTGCATAGT
GGTGACTCAATCGCTGTATATCCACCACAAACTTTGACAGAAGACGAGTTAGCAACACTT
GAGGACTATACTATAAAATTAGCTAAAGGTTTAAACATCATTGGCTTAATCAACATTCAA
TTCGTTATAGCTCACGATGGTGTGTATGTTTTAGAAGTAAATCCACGTTCTAGTAGAACG
GTACCATTCTTAAGTAAAATTACTGATATTCCAATGGCACAATTAGCTATGCGAGCAATC
ATTGGGGAAAAACTAACAGATATGGGTTATCAAGAAGGGGTTCAACCATATGCTGAGGGT
GTCTTTGTGAAAGCACCAGTATTTAGTTTTAATAAATTGAAAAATGTTGATATTACTTTA
GGACCTGAAATGAAGTCAACAGGTGAAGTGATGGGGAAAGATACTACATTAGAAAAGGCG
TTATTCAAAGGGTTAACAGGTAGTGGCGTTGAAGTTAAAGATCACGGTACAGTATTAATG
ACCGTCAGTGACAAAGATAAAGAGGAAGTTGTTAAATTGGCACAACGCTTAAATGAAGTT
GGCTATAAAATTTTAGCAACGTCTGGAACAGCTAATAAATTAGCTGAGTATGACATACCT
GCAGAAGTAGTAGGCAAAATTGGTGGCGAAAATGATTTATTAACACGTATTCAAAATGGT
GATGTTCAAATCGTTATAAATACAATGACTAAAGGTAAAGAAGTAGAAAGGGATGGCTTC
CAAATTAGACGTACTACAGTTGAAAATGGTATTCCATGTTTGACATCTTTAGATACAGCT
AATGCCTTAACGAATGTAATTGAAAGTATGACATTTACAATGCGTCAAATGTAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1260
1320
1380
1440
1500
1560
1620
1680
1740
1800
1860
1920
1980
2040
2100
2160
2220
2280
2340
2400
2460
2520
2580
2640
2700
2760
2820
2880
2940
3000
3060
3120
3174
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_01170
- symbol: CarB
- description: carbamoyl phosphate synthase large subunit
- length: 1057
- theoretical pI: 4.60041
- theoretical MW: 117185
- GRAVY: -0.194229
⊟Function[edit | edit source]
- reaction: EC 6.3.5.5? ExPASyCarbamoyl-phosphate synthase (glutamine-hydrolyzing) 2 ATP + L-glutamine + HCO3(-) + H2O = 2 ADP + phosphate + L-glutamate + carbamoyl phosphate
- TIGRFAM: Purines, pyrimidines, nucleosides, and nucleotides Pyrimidine ribonucleotide biosynthesis carbamoyl-phosphate synthase, large subunit (TIGR01369; EC 6.3.5.5; HMM-score: 1581.7)and 15 moreEnergy metabolism Glycolysis/gluconeogenesis pyruvate carboxylase (TIGR01235; EC 6.4.1.1; HMM-score: 128)Fatty acid and phospholipid metabolism Biosynthesis acetyl-CoA carboxylase, biotin carboxylase subunit (TIGR00514; EC 6.3.4.14; HMM-score: 99.8)Purines, pyrimidines, nucleosides, and nucleotides Purine ribonucleotide biosynthesis phosphoribosylglycinamide formyltransferase 2 (TIGR01142; EC 2.1.2.-; HMM-score: 96.2)Cell envelope Biosynthesis and degradation of murein sacculus and peptidoglycan D-alanine--D-alanine ligase (TIGR01205; EC 6.3.2.4; HMM-score: 91.7)Central intermediary metabolism Nitrogen metabolism urea carboxylase (TIGR02712; EC 6.3.4.6; HMM-score: 81.9)Purines, pyrimidines, nucleosides, and nucleotides Purine ribonucleotide biosynthesis phosphoribosylamine--glycine ligase (TIGR00877; EC 6.3.4.13; HMM-score: 74.7)lysine biosynthesis enzyme LysX (TIGR02144; HMM-score: 68.1)Cellular processes Biosynthesis of natural products cyanophycin synthetase (TIGR02068; EC 6.3.2.29,6.3.2.30; HMM-score: 67.6)Amino acid biosynthesis Other pyrrolysine biosynthesis protein PylC (TIGR03909; HMM-score: 66.4)alpha-L-glutamate ligase, RimK family (TIGR00768; EC 6.3.2.-; HMM-score: 57.8)Purines, pyrimidines, nucleosides, and nucleotides Purine ribonucleotide biosynthesis phosphoribosylaminoimidazole carboxylase, ATPase subunit (TIGR01161; EC 4.1.1.21; HMM-score: 52.1)Biosynthesis of cofactors, prosthetic groups, and carriers Other coenzyme gamma-F420-2:alpha-L-glutamate ligase (TIGR04443; EC 6.3.2.32; HMM-score: 35.6)ATP-grasp enzyme, GAK system (TIGR04356; HMM-score: 27.1)Purines, pyrimidines, nucleosides, and nucleotides Purine ribonucleotide biosynthesis phosphoribosylaminoimidazolecarboxamide formyltransferase/IMP cyclohydrolase (TIGR00355; EC 2.1.2.3,3.5.4.10; HMM-score: 21.7)Biosynthesis of cofactors, prosthetic groups, and carriers Heme, porphyrin, and cobalamin putative heme d1 biosynthesis radical SAM protein NirJ2 (TIGR04055; EC 1.3.99.-; HMM-score: 13.8)
- TheSEED :
- Carbamoyl-phosphate synthase large chain (EC 6.3.5.5)
- PFAM: ATP-grasp (CL0179) CPSase_L_D2; Carbamoyl-phosphate synthase L chain, ATP binding domain (PF02786; HMM-score: 407.3)and 14 moreHTH (CL0123) CPSase_L_D3; Carbamoyl-phosphate synthetase large chain, oligomerisation domain (PF02787; HMM-score: 92.1)no clan defined MGS; MGS-like domain (PF02142; HMM-score: 82.1)ATP-grasp (CL0179) Dala_Dala_lig_C; D-ala D-ala ligase C-terminus (PF07478; HMM-score: 72.8)ATP-grasp_3; ATP-grasp domain (PF02655; HMM-score: 70.8)ATP-grasp; ATP-grasp domain (PF02222; HMM-score: 66.2)GARS_A; Phosphoribosylglycinamide synthetase, ATP-grasp (A) domain (PF01071; HMM-score: 55.7)ATPgrasp_Ter; ATP-grasp in the biosynthetic pathway with Ter operon (PF15632; HMM-score: 54.6)ATP-grasp_4; ATP-grasp domain (PF13535; HMM-score: 34.4)ATP-grasp_5; ATP-grasp domain (PF13549; HMM-score: 32.9)RimK; RimK-like ATP-grasp domain (PF08443; HMM-score: 30.4)ATP-grasp_2; ATP-grasp domain (PF08442; HMM-score: 24.4)NADP_Rossmann (CL0063) ADH_zinc_N; Zinc-binding dehydrogenase (PF00107; HMM-score: 14.6)CoA_binding_2; CoA binding domain (PF13380; HMM-score: 13.2)no clan defined PrpR_N; Propionate catabolism activator (PF06506; HMM-score: 12.7)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors: Mg2+, Mn2+
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.97
- Cytoplasmic Membrane Score: 0
- Cellwall Score: 0.01
- Extracellular Score: 0.02
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.8076
- Cytoplasmic Membrane Score: 0.1737
- Cell wall & surface Score: 0.0041
- Extracellular Score: 0.0146
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.152665
- TAT(Tat/SPI): 0.008979
- LIPO(Sec/SPII): 0.022409
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MPKRNDIKTILVIGSGPIIIGQAAEFDYAGTQACLALKEEGYRVILVNSNPATIMTDKEIADKVYIEPLTHDFIARIIRKEQPDALLPTLGGQTGLNMAIQLHESGVLQDNNVQLLGTELTSIQQAEDREMFRTLMNDLNVPVPESDIVNTVEQAFKFKEQVGYPLIVRPAFTMGGTGGGICHNDEELHEIVSNGLHYSPATQCLLEKSIAGFKEIEYEVMRDKNDNAIVVCNMENIDPVGIHTGDSIVVAPSQTLSDVEYQMLRDVSLKVIRALGIEGGCNVQLALDPHSFDYYIIEVNPRVSRSSALASKATGYPIAKLAAKIAVGLTLDEMLNPITGTSYAAFEPTLDYVISKIPRFPFDKFEKGERELGTQMKATGEVMAIGRTYEESLLKAIRSLEYGVHHLGLPNGESFDLDYIKERISHQDDERLFFIGEAIRRGTTLEEIHNMTQIDYFFLHKFQNIIDIEHQLKEHQGDLEYLKYAKDYGFSDKTIAHRFNMTEEEVYQLRMENDIKPVYKMVDTCAAEFESSTPYYYGTYETENESIVTDKEKILVLGSGPIRIGQGVEFDYATVHAVWAIQKAGYEAIIVNNNPETVSTDFSISDKLYFEPLTEEDVMNIINLEKPKGVVVQFGGQTAINLADKLAKHGVKILGTSLENLNRAEDRKEFEALLRKINVPQPQGKTATSPEEALANAAEIGYPVVVRPSYVLGGRAMEIVDNDKELENYMTQAVKASPEHPVLVDRYLTGKEIEVDAICDGETVIIPGIMEHIERAGVHSGDSIAVYPPQTLTEDELATLEDYTIKLAKGLNIIGLINIQFVIAHDGVYVLEVNPRSSRTVPFLSKITDIPMAQLAMRAIIGEKLTDMGYQEGVQPYAEGVFVKAPVFSFNKLKNVDITLGPEMKSTGEVMGKDTTLEKALFKGLTGSGVEVKDHGTVLMTVSDKDKEEVVKLAQRLNEVGYKILATSGTANKLAEYDIPAEVVGKIGGENDLLTRIQNGDVQIVINTMTKGKEVERDGFQIRRTTVENGIPCLTSLDTANALTNVIESMTFTMRQM
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas [1] [2]
- protein localization: data available for COL
- quantitative data / protein copy number per cell: data available for COL
- interaction partners:
SAOUHSC_01232 (rpsB) 30S ribosomal protein S2 [3] (data from MRSA252) SAOUHSC_02494 (rpsE) 30S ribosomal protein S5 [3] (data from MRSA252) SAOUHSC_00530 elongation factor Tu [3] (data from MRSA252) SAOUHSC_00679 hypothetical protein [3] (data from MRSA252) SAOUHSC_01040 pyruvate dehydrogenase complex, E1 component subunit alpha [3] (data from MRSA252) SAOUHSC_01819 hypothetical protein [3] (data from MRSA252)
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: SAOUHSC_01164 > SAOUHSC_01165 > pyrB > pyrC > SAOUHSC_01169 > carB > SAOUHSC_01171 > pyrEpredicted SigA promoter [4] : lspA > SAOUHSC_01163 > S480 > S481 > SAOUHSC_01164 > S482 > SAOUHSC_01165 > pyrB > pyrC > SAOUHSC_01169 > carB > S483 > SAOUHSC_01171 > pyrE > SAOUHSC_01173 > S484 > SAOUHSC_01174 > S485predicted SigA promoter [4] : S481 > SAOUHSC_01164 > S482 > SAOUHSC_01165 > pyrB > pyrC > SAOUHSC_01169 > carB > S483 > SAOUHSC_01171 > pyrE > SAOUHSC_01173 > S484 > SAOUHSC_01174 > S485
⊟Regulation[edit | edit source]
- regulator: pyrR leader (transcription antitermination) regulon
pyrR leader (5' cis-acting region) important in Pyrimidine metabolism; transcription unit transferred from N315 data RegPrecise
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [4]
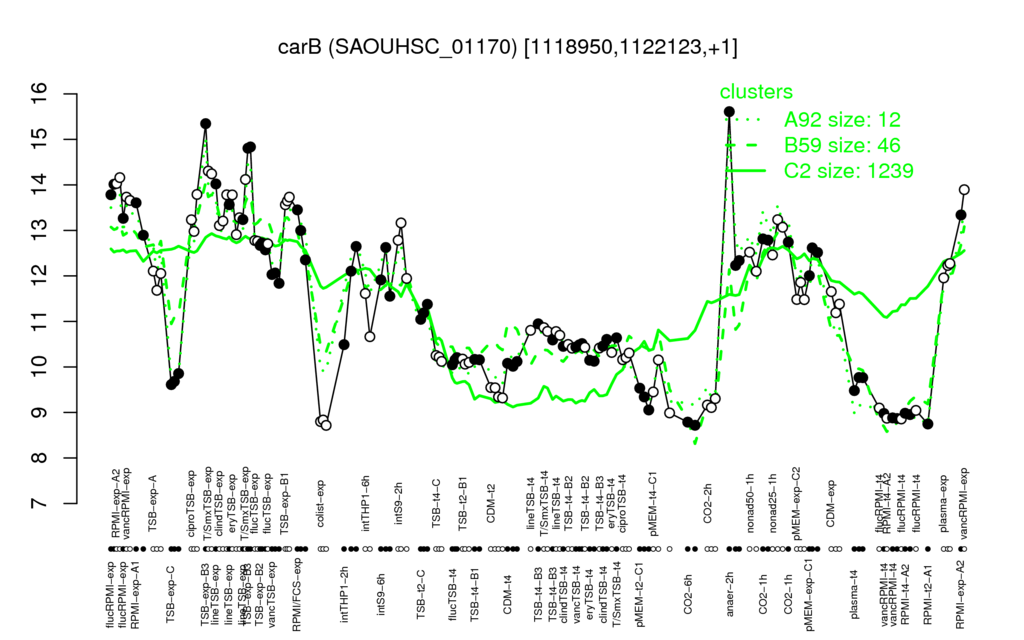 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Maren Depke, Stephan Michalik, Alexander Rabe, Kristin Surmann, Lars Brinkmann, Nico Jehmlich, Jörg Bernhardt, Michael Hecker, Bernd Wollscheid, Zhi Sun, Robert L Moritz, Uwe Völker, Frank Schmidt
A peptide resource for the analysis of Staphylococcus aureus in host-pathogen interaction studies.
Proteomics: 2015, 15(21);3648-61
[PubMed:26224020] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Maren Depke, Annette Murr, Manuela Gesell Salazar, Ulrike Kusebauch, Zhi Sun, Tanja C Meyer, Kristin Surmann, Henrike Pförtner, Petra Hildebrandt, Stefan Weiss, Laura Marcela Palma Medina, Melanie Gutjahr, Elke Hammer, Dörte Becher, Thomas Pribyl, Sven Hammerschmidt, Eric W Deutsch, Samuel L Bader, Michael Hecker, Robert L Moritz, Ulrike Mäder, Uwe Völker, Frank Schmidt
A global Staphylococcus aureus proteome resource applied to the in vivo characterization of host-pathogen interactions.
Sci Rep: 2017, 7(1);9718
[PubMed:28887440] [WorldCat.org] [DOI] (I e) - ↑ 3.0 3.1 3.2 3.3 3.4 3.5 Artem Cherkasov, Michael Hsing, Roya Zoraghi, Leonard J Foster, Raymond H See, Nikolay Stoynov, Jihong Jiang, Sukhbir Kaur, Tian Lian, Linda Jackson, Huansheng Gong, Rick Swayze, Emily Amandoron, Farhad Hormozdiari, Phuong Dao, Cenk Sahinalp, Osvaldo Santos-Filho, Peter Axerio-Cilies, Kendall Byler, William R McMaster, Robert C Brunham, B Brett Finlay, Neil E Reiner
Mapping the protein interaction network in methicillin-resistant Staphylococcus aureus.
J Proteome Res: 2011, 10(3);1139-50
[PubMed:21166474] [WorldCat.org] [DOI] (I p) - ↑ 4.0 4.1 4.2 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
