⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_02536
- pan locus tag?: SAUPAN005729000
- symbol: SAOUHSC_02536
- pan gene symbol?: moaA
- synonym:
- product: molybdenum cofactor biosynthesis protein A
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_02536
- symbol: SAOUHSC_02536
- product: molybdenum cofactor biosynthesis protein A
- replicon: chromosome
- strand: -
- coordinates: 2336046..2337068
- length: 1023
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3921126 NCBI
- RefSeq: YP_501000 NCBI
- BioCyc: G1I0R-2397 BioCyc
- MicrobesOnline: 1290971 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021ATGGTAGAACAAATAAAAGATAAACTAGGACGTCCCATCCGTGACTTACGGTTATCTGTG
ACAGATCGGTGTAACTTTAGGTGTGATTATTGCATGCCTAAAGAGGTATTTGGAGATGAT
TTCGTATTTTTACCTAAAAATGAACTTTTAACGTTTGATGAAATGGCTAGAATCGCTAAG
GTATATGCAGAATTAGGTGTAAAAAAAATACGCATTACAGGTGGAGAACCATTGATGCGA
CGGGATTTAGATGTACTTATAGCTAAATTAAATCAAATCGATGGTATTGAAGATATTGGT
TTGACTACAAATGGTTTGTTATTAAAAAAGCATGGACAAAAGTTATATGATGCTGGGCTG
CGCAGAATTAATGTCAGTTTGGATGCTATTGATGATACGCTATTTCAATCAATCAATAAT
CGTAATATTAAAGCGACTACGATTTTAGAACAAATTGATTACGCGACGTCTATTGGTTTG
AATGTAAAAGTAAATGTTGTTATACAAAAAGGTATTAACGATGATCAAATCATACCAATG
CTTGAATATTTTAAAGATAAACATATAGAGATTCGATTTATAGAATTTATGGATGTTGGT
AATGATAATGGATGGGATTTCAGTAAAGTTGTAACTAAAGATGAAATGCTTACAATGATA
GAGCAGCACTTTGAAATCGATCCTGTAGAACCAAAATATTTTGGGGAAGTAGCAAAATAT
TATCGCCATAAGGATAATGGTGTTCAATTTGGTTTGATTACAAGTGTTTCACAATCATTT
TGTTCTACATGTACACGCGCAAGGCTGTCATCAGATGGGAAGTTTTACGGATGTTTATTT
GCAACTGTCGATGGATTTAACGTTAAAGCGTTTATTCGTTCTGGCGTGACCGACGAAGAA
TTAAAAGAACAATTTAAAGCTTTATGGCAAATAAGAGATGATCGATATTCAGATGAGAGA
ACTGCTCAAACAGTTGCCAATCGTCAACGTAAAAAGATAAACATGAATTATATTGGTGGT
TAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1023
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_02536
- symbol: SAOUHSC_02536
- description: molybdenum cofactor biosynthesis protein A
- length: 340
- theoretical pI: 6.53074
- theoretical MW: 39077.6
- GRAVY: -0.343529
⊟Function[edit | edit source]
- reaction: EC 4.1.99.22? ExPASyGTP 3',8-cyclase GTP + S-adenosyl-L-methionine + reduced electron acceptor = (8S)-3',8-cyclo-7,8-dihydroguanosine 5'-triphosphate + 5'-deoxyadenosine + L-methionine + oxidized electron acceptor
- TIGRFAM: Biosynthesis of cofactors, prosthetic groups, and carriers Molybdopterin molybdenum cofactor biosynthesis protein A (TIGR02666; HMM-score: 402.8)and 63 moreBiosynthesis of cofactors, prosthetic groups, and carriers Molybdopterin probable molybdenum cofactor biosynthesis protein A (TIGR02668; HMM-score: 233.3)Biosynthesis of cofactors, prosthetic groups, and carriers Other coenzyme PQQ biosynthesis enzyme PqqE (TIGR02109; HMM-score: 61.3)tungsten cofactor oxidoreducase radical SAM maturase (TIGR04317; HMM-score: 59.1)pseudo-rSAM protein/SPASM domain protein (TIGR04347; HMM-score: 52.2)putative metalloenzyme radical SAM/SPASM domain maturase (TIGR04311; HMM-score: 51.9)Biosynthesis of cofactors, prosthetic groups, and carriers Heme, porphyrin, and cobalamin heme b synthase (TIGR04545; EC 1.3.99.-; HMM-score: 48.9)Biosynthesis of cofactors, prosthetic groups, and carriers Heme, porphyrin, and cobalamin putative heme d1 biosynthesis radical SAM protein NirJ2 (TIGR04055; EC 1.3.99.-; HMM-score: 46.8)Biosynthesis of cofactors, prosthetic groups, and carriers Heme, porphyrin, and cobalamin 12,18-didecarboxysiroheme deacetylase (TIGR04546; HMM-score: 46.7)radical SAM enzyme, rSAM/lipoprotein system (TIGR04133; HMM-score: 44.9)GeoRSP system radical SAM/SPASM protein (TIGR04303; HMM-score: 43.1)Biosynthesis of cofactors, prosthetic groups, and carriers Heme, porphyrin, and cobalamin heme d1 biosynthesis radical SAM protein NirJ (TIGR04051; HMM-score: 41.8)Cellular processes Toxin production and resistance cytosylglucuronate decarboxylase (TIGR04466; EC 4.1.-.-; HMM-score: 41.8)Unknown function Enzymes of unknown specificity radical SAM protein, BA_1875 family (TIGR04053; HMM-score: 41.2)Biosynthesis of cofactors, prosthetic groups, and carriers Heme, porphyrin, and cobalamin putative heme d1 biosynthesis radical SAM protein NirJ1 (TIGR04054; HMM-score: 41)hopanoid biosynthesis associated radical SAM protein HpnH (TIGR03470; HMM-score: 40.6)SynChlorMet cassette radical SAM/SPASM protein ScmF (TIGR04251; HMM-score: 35.5)peptide-modifying radical SAM enzyme CbpB (TIGR04163; HMM-score: 34)radical SAM protein, TatD family-associated (TIGR04038; HMM-score: 33.7)radical SAM peptide maturase, GG-Bacteroidales family (TIGR04148; HMM-score: 33.1)SynChlorMet cassette radical SAM/SPASM protein ScmE (TIGR04250; HMM-score: 33)Unknown function Enzymes of unknown specificity archaeal radical SAM protein, PTO1314 family (TIGR03961; HMM-score: 32.8)antiviral radical SAM protein viperin (TIGR04278; HMM-score: 32.4)Protein synthesis tRNA and rRNA base modification putative 7-cyano-7-deazaguanosine (preQ0) biosynthesis protein QueE (TIGR03963; HMM-score: 32.3)nif11-class peptide radical SAM maturase 3 (TIGR04103; HMM-score: 31.9)Unknown function Enzymes of unknown specificity Y_X(10)_GDL-associated radical SAM protein (TIGR03913; HMM-score: 30.1)His-Xaa-Ser system radical SAM maturase HxsC (TIGR03977; HMM-score: 30.1)radical SAM/SPASM domain protein, FxsB family (TIGR04269; HMM-score: 30.1)Unknown function Enzymes of unknown specificity mycofactocin radical SAM maturase (TIGR03962; HMM-score: 29.8)arginine 2,3-aminomutase (TIGR04468; EC 5.4.3.-; HMM-score: 29.7)Protein fate Protein modification and repair anaerobic ribonucleoside-triphosphate reductase activating protein (TIGR02495; EC 1.97.-.-; HMM-score: 29)Purines, pyrimidines, nucleosides, and nucleotides 2'-Deoxyribonucleotide metabolism anaerobic ribonucleoside-triphosphate reductase activating protein (TIGR02495; EC 1.97.-.-; HMM-score: 29)Biosynthesis of cofactors, prosthetic groups, and carriers Biotin biotin synthase (TIGR00433; EC 2.8.1.6; HMM-score: 26.4)Biosynthesis of cofactors, prosthetic groups, and carriers Other nitrogenase cofactor biosynthesis protein NifB (TIGR01290; HMM-score: 26.3)Central intermediary metabolism Nitrogen fixation nitrogenase cofactor biosynthesis protein NifB (TIGR01290; HMM-score: 26.3)radical SAM/Cys-rich domain protein (TIGR04167; HMM-score: 26.3)Cellular processes Toxin production and resistance Cys-rich peptide radical SAM maturase CcpM (TIGR04068; HMM-score: 25.5)sporulation killing factor system radical SAM maturase (TIGR04403; HMM-score: 24.5)Protein fate Protein modification and repair KxxxW cyclic peptide radical SAM maturase (TIGR04080; HMM-score: 24.4)Cellular processes Adaptations to atypical conditions KamA family protein (TIGR00238; HMM-score: 23.7)Cellular processes Biosynthesis of natural products SCIFF radical SAM maturase (TIGR03974; HMM-score: 23.6)radical SAM/SPASM domain protein, ACGX system (TIGR04340; HMM-score: 23.6)pseudo-rSAM protein, GG-Bacteroidales system (TIGR04150; HMM-score: 21.4)lysine-2,3-aminomutase-related protein (TIGR03822; EC 5.4.3.-; HMM-score: 21)radical SAM enzyme, TIGR04100 family (TIGR04100; HMM-score: 20.6)putative peptide-modifying radical SAM enzyme, AF0577 family (TIGR04084; HMM-score: 20.2)putative 7-cyano-7-deazaguanosine (preQ0) biosynthesis protein QueE (TIGR04349; HMM-score: 18.9)Protein synthesis tRNA and rRNA base modification 7-cyano-7-deazaguanosine (preQ0) biosynthesis protein QueE (TIGR03365; HMM-score: 18.7)Unknown function Enzymes of unknown specificity radical SAM/CxCxxxxC motif protein YfkAB (TIGR04478; HMM-score: 18.1)Protein synthesis tRNA and rRNA base modification wyosine biosynthesis protein TYW1 (TIGR03972; HMM-score: 17.6)radical SAM/SPASM domain peptide maturase, XyeB family (TIGR04496; HMM-score: 17.2)Protein fate Protein modification and repair EF-P beta-lysylation protein EpmB (TIGR03821; EC 5.4.3.-; HMM-score: 16.8)lysine-2,3-aminomutase (TIGR03820; EC 5.4.3.2; HMM-score: 16.6)putative radical SAM enzyme, TIGR03279 family (TIGR03279; HMM-score: 16.5)Protein fate Protein modification and repair anaerobic sulfatase maturase (TIGR03942; EC 1.1.99.-; HMM-score: 16)Cellular processes Toxin production and resistance nif11-like peptide radical SAM maturase (TIGR04064; HMM-score: 15.6)DNA metabolism DNA replication, recombination, and repair DNA repair protein rad18 (TIGR00599; HMM-score: 15.4)Protein fate Protein modification and repair quinohemoprotein amine dehydrogenase maturation protein (TIGR03906; HMM-score: 14.1)radical SAM/SPASM domain protein maturase (TIGR04463; HMM-score: 14)His-Xaa-Ser system radical SAM maturase HxsB (TIGR03978; HMM-score: 13.4)Biosynthesis of cofactors, prosthetic groups, and carriers Other 7,8-didemethyl-8-hydroxy-5-deazariboflavin synthase, CofH subunit (TIGR03551; EC 2.5.1.77; HMM-score: 12)Biosynthesis of cofactors, prosthetic groups, and carriers Other 7,8-didemethyl-8-hydroxy-5-deazariboflavin synthase, CofG subunit (TIGR03550; EC 2.5.1.77; HMM-score: 11.3)Biosynthesis of cofactors, prosthetic groups, and carriers Other homocitrate synthase (TIGR02660; EC 2.3.3.14; HMM-score: 11)Central intermediary metabolism Nitrogen fixation homocitrate synthase (TIGR02660; EC 2.3.3.14; HMM-score: 11)
- TheSEED :
- GTP 3',8-cyclase (EC 4.1.99.22)
- PFAM: no clan defined Mob_synth_C; Molybdenum Cofactor Synthesis C (PF06463; HMM-score: 133.7)TIM_barrel (CL0036) Radical_SAM; Radical SAM superfamily (PF04055; HMM-score: 114.7)and 5 moreRadical_SAM_2; Radical SAM-like domain (PF19238; HMM-score: 23)4Fe-4S (CL0344) Fer4_12; 4Fe-4S single cluster domain (PF13353; HMM-score: 22.9)no clan defined Nif11; Nif11 domain (PF07862; HMM-score: 14.8)DUF3194; Protein of unknown function (DUF3194) (PF11419; HMM-score: 13.3)Fe_hyd_lg_C; Iron only hydrogenase large subunit, C-terminal domain (PF02906; HMM-score: 13)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors: [4Fe-4S] cluster
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 7.5
- Cytoplasmic Membrane Score: 1.15
- Cellwall Score: 0.62
- Extracellular Score: 0.73
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.9749
- Cytoplasmic Membrane Score: 0.0022
- Cell wall & surface Score: 0
- Extracellular Score: 0.0229
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.007959
- TAT(Tat/SPI): 0.000492
- LIPO(Sec/SPII): 0.001254
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MVEQIKDKLGRPIRDLRLSVTDRCNFRCDYCMPKEVFGDDFVFLPKNELLTFDEMARIAKVYAELGVKKIRITGGEPLMRRDLDVLIAKLNQIDGIEDIGLTTNGLLLKKHGQKLYDAGLRRINVSLDAIDDTLFQSINNRNIKATTILEQIDYATSIGLNVKVNVVIQKGINDDQIIPMLEYFKDKHIEIRFIEFMDVGNDNGWDFSKVVTKDEMLTMIEQHFEIDPVEPKYFGEVAKYYRHKDNGVQFGLITSVSQSFCSTCTRARLSSDGKFYGCLFATVDGFNVKAFIRSGVTDEELKEQFKALWQIRDDRYSDERTAQTVANRQRKKINMNYIGG
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas [1] [2]
- protein localization: data available for COL
- quantitative data / protein copy number per cell:
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: SAOUHSC_02536 < mobA < SAOUHSC_02538 < SAOUHSC_02540 < SAOUHSC_02541 < SAOUHSC_02542predicted SigA promoter [3] : SAOUHSC_02532 < S978 < SAOUHSC_02534 < SAOUHSC_02535 < S979 < SAOUHSC_02536 < mobA < SAOUHSC_02538 < SAOUHSC_02540 < SAOUHSC_02541 < SAOUHSC_02542 < S980 < SAOUHSC_02544 < SAOUHSC_02545 < S981 < SAOUHSC_02546 < SAOUHSC_02547 < SAOUHSC_02549
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [3]
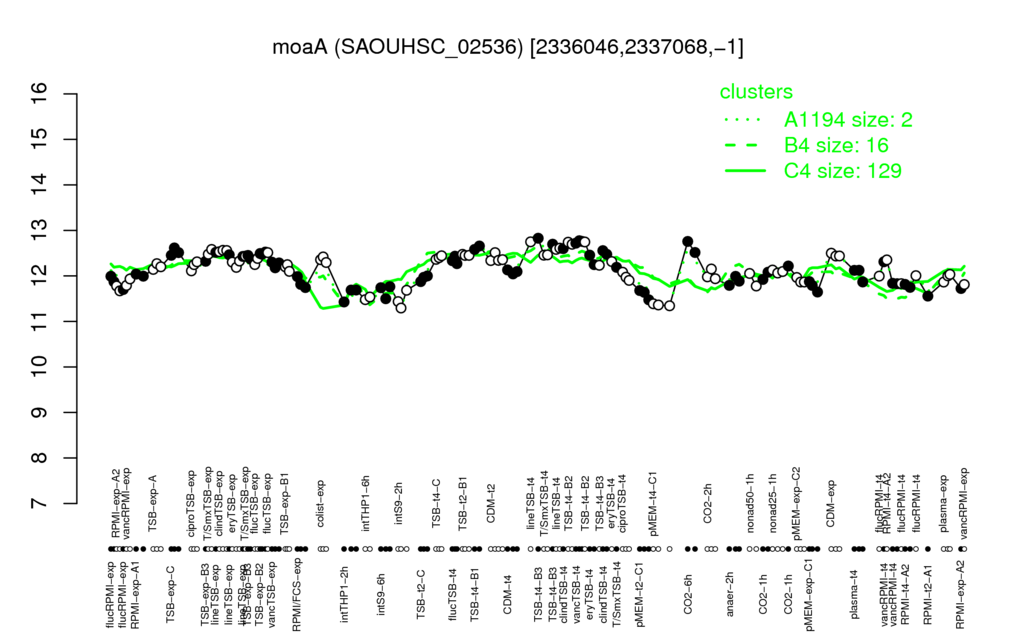 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Maren Depke, Stephan Michalik, Alexander Rabe, Kristin Surmann, Lars Brinkmann, Nico Jehmlich, Jörg Bernhardt, Michael Hecker, Bernd Wollscheid, Zhi Sun, Robert L Moritz, Uwe Völker, Frank Schmidt
A peptide resource for the analysis of Staphylococcus aureus in host-pathogen interaction studies.
Proteomics: 2015, 15(21);3648-61
[PubMed:26224020] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Maren Depke, Annette Murr, Manuela Gesell Salazar, Ulrike Kusebauch, Zhi Sun, Tanja C Meyer, Kristin Surmann, Henrike Pförtner, Petra Hildebrandt, Stefan Weiss, Laura Marcela Palma Medina, Melanie Gutjahr, Elke Hammer, Dörte Becher, Thomas Pribyl, Sven Hammerschmidt, Eric W Deutsch, Samuel L Bader, Michael Hecker, Robert L Moritz, Ulrike Mäder, Uwe Völker, Frank Schmidt
A global Staphylococcus aureus proteome resource applied to the in vivo characterization of host-pathogen interactions.
Sci Rep: 2017, 7(1);9718
[PubMed:28887440] [WorldCat.org] [DOI] (I e) - ↑ 3.0 3.1 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
⊟Relevant publications[edit | edit source]
Petra Hänzelmann, Hermann Schindelin
Crystal structure of the S-adenosylmethionine-dependent enzyme MoaA and its implications for molybdenum cofactor deficiency in humans.
Proc Natl Acad Sci U S A: 2004, 101(35);12870-5
[PubMed:15317939] [WorldCat.org] [DOI] (P p)Petra Hänzelmann, Hermann Schindelin
Binding of 5'-GTP to the C-terminal FeS cluster of the radical S-adenosylmethionine enzyme MoaA provides insights into its mechanism.
Proc Natl Acad Sci U S A: 2006, 103(18);6829-34
[PubMed:16632608] [WorldCat.org] [DOI] (P p)Nicholas S Lees, Petra Hänzelmann, Heather L Hernandez, Sowmya Subramanian, Hermann Schindelin, Michael K Johnson, Brian M Hoffman
ENDOR spectroscopy shows that guanine N1 binds to [4Fe-4S] cluster II of the S-adenosylmethionine-dependent enzyme MoaA: mechanistic implications.
J Am Chem Soc: 2009, 131(26);9184-5
[PubMed:19566093] [WorldCat.org] [DOI] (I p)
