Jump to navigation
Jump to search
NCBI: 10-JUN-2013
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus COL
- locus tag: SACOL1520 [new locus tag: SACOL_RS07750 ]
- pan locus tag?: SAUPAN003944000
- symbol: SACOL1520
- pan gene symbol?: bdr
- synonym: ypdA
- product: pyridine nucleotide-disulfide oxidoreductase
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SACOL1520 [new locus tag: SACOL_RS07750 ]
- symbol: SACOL1520
- product: pyridine nucleotide-disulfide oxidoreductase
- replicon: chromosome
- strand: -
- coordinates: 1558965..1559951
- length: 987
- essential: unknown other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3237980 NCBI
- RefSeq: YP_186363 NCBI
- BioCyc: see SACOL_RS07750
- MicrobesOnline: 912971 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961ATGCAAAAAGTTGAAAGTATCATAATTGGTGGAGGGCCATGCGGATTAAGTGCGGCTATT
GAACAAAAAAGAAAAGGTATTGATACCTTAATTATTGAAAAGGGTAATGTCGTTGAATCA
ATCTACAATTATCCTACTCACCAAACATTTTTCTCATCAAGTGATAAATTAAGTATTGGG
GACGTACCGTTTATCGTTGAAGAAAGTAAACCAAGACGTAATCAAGCGCTAGTTTATTAC
CGAGAAGTTGTAAAACATCATCAATTAAAAGTAAATGCATTTGAAGAAGTATTAACTGTT
AAAAAAATGAATAATAAATTTACTATTACTACGACGAAAGATGTTTATGAATGTCGATTT
TTAACAATCGCGACAGGCTATTATGGTCAGCATAATACATTAGAAGTTGAAGGTGCGGAT
TTACCTAAAGTGTTCCATTATTTTAAAGAGGCACATCCGTATTTTGATCAAGATGTTGTA
ATTATCGGTGGTAAGAATTCGGCTATCGATGCTGCTTTGGAGTTGGAAAAAGCTGGTGCT
AACGTGACGGTTCTATATCGTGGTGGAGATTATTCGCCTTCAATTAAACCGTGGATACTT
CCAAATTTCACAGCATTAGTAAATCATGAAAAAATTGACATGGAATTTAATGCTAATGTT
ACCCAAATAACTGAAGATACTGTGACTTATGAAGTAAATGGTGAAAGTAAAACGATACAC
AATGATTATGTATTTGCGATGATTGGTTATCATCCCGATTATGAATTTTTAAAATCTGTA
GGCATTCAAATTAATACAAATGAATTTGGAACAGCGCCTATGTATAATAAAGAAACATAC
GAAACAAATATCGAAAATTGCTATATTGCAGGTGTAATTGCTGCAGGGAACGATGCGAAT
ACCATTTTTATTGAAAATGGTAAATTCCACGGGGGCATTATTGCTCAAAGCATGCTAGCT
AAGAAACAAACGCCCTTAGAATCATAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
987
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SACOL1520 [new locus tag: SACOL_RS07750 ]
- symbol: SACOL1520
- description: pyridine nucleotide-disulfide oxidoreductase
- length: 328
- theoretical pI: 5.39888
- theoretical MW: 36736.3
- GRAVY: -0.271646
⊟Function[edit | edit source]
- TIGRFAM: Unknown function Enzymes of unknown specificity putative bacillithiol system oxidoreductase, YpdA family (TIGR04018; EC 1.8.-.-; HMM-score: 457.2)and 42 moreEnergy metabolism Electron transport thioredoxin-disulfide reductase (TIGR01292; EC 1.8.1.9; HMM-score: 87.3)Cellular processes Detoxification CoA-disulfide reductase (TIGR03385; EC 1.8.1.14; HMM-score: 61.7)dihydrolipoyl dehydrogenase (TIGR01350; EC 1.8.1.4; HMM-score: 59.3)Cellular processes Detoxification alkyl hydroperoxide reductase subunit F (TIGR03140; EC 1.8.1.-; HMM-score: 50.8)Cellular processes Adaptations to atypical conditions alkyl hydroperoxide reductase subunit F (TIGR03140; EC 1.8.1.-; HMM-score: 50.8)Cellular processes Detoxification mercury(II) reductase (TIGR02053; EC 1.16.1.1; HMM-score: 49.4)putative selenate reductase, YgfK subunit (TIGR03315; HMM-score: 48.8)putative alkyl hydroperoxide reductase F subunit (TIGR03143; EC 1.6.4.-; HMM-score: 48.2)Amino acid biosynthesis Glutamate family glutamate synthase (NADPH), homotetrameric (TIGR01316; EC 1.4.1.13; HMM-score: 38.6)flavin-dependent oxidoreductase, MSMEG_0569 family (TIGR04046; HMM-score: 35.1)Unknown function Enzymes of unknown specificity flavoprotein, HI0933 family (TIGR00275; HMM-score: 34.1)Energy metabolism Electron transport glutathione-disulfide reductase (TIGR01421; EC 1.8.1.7; HMM-score: 29.5)squalene-associated FAD-dependent desaturase (TIGR03467; HMM-score: 25.6)Biosynthesis of cofactors, prosthetic groups, and carriers Thiamine thiazole biosynthesis enzyme (TIGR00292; HMM-score: 25.5)Biosynthesis of cofactors, prosthetic groups, and carriers Chlorophyll and bacteriochlorphyll geranylgeranyl reductase family (TIGR02032; EC 1.3.1.-; HMM-score: 25.2)Biosynthesis of cofactors, prosthetic groups, and carriers Chlorophyll and bacteriochlorphyll geranylgeranyl reductase (TIGR02023; EC 1.3.1.-; HMM-score: 23.3)Energy metabolism Other 4-hydroxybenzoate 3-monooxygenase (TIGR02360; EC 1.14.13.2; HMM-score: 20.8)Biosynthesis of cofactors, prosthetic groups, and carriers Heme, porphyrin, and cobalamin precorrin 3B synthase CobZ (TIGR02485; HMM-score: 20.3)pyridine nucleotide-disulfide oxidoreductase family protein (TIGR03169; HMM-score: 20.2)mycothione reductase (TIGR03452; EC 1.8.1.15; HMM-score: 19.2)glutamate synthase, small subunit (TIGR01318; HMM-score: 19)Central intermediary metabolism Nitrogen metabolism nitrite reductase [NAD(P)H], large subunit (TIGR02374; EC 1.7.1.4; HMM-score: 18.4)glutamate synthase, NADH/NADPH, small subunit (TIGR01317; EC 1.4.1.-; HMM-score: 18.2)Energy metabolism Anaerobic glycerol-3-phosphate dehydrogenase, anaerobic, B subunit (TIGR03378; EC 1.1.5.3; HMM-score: 18.1)Energy metabolism Amino acids and amines sarcosine oxidase, alpha subunit family (TIGR01372; HMM-score: 17.1)Energy metabolism Amino acids and amines sarcosine oxidase, monomeric form (TIGR01377; HMM-score: 16.8)mycofactocin system FadH/OYE family oxidoreductase 2 (TIGR03997; EC 1.-.-.-; HMM-score: 16.8)Biosynthesis of cofactors, prosthetic groups, and carriers Other phytoene desaturase (TIGR02734; EC 1.14.99.-; HMM-score: 16.7)Energy metabolism TCA cycle succinate dehydrogenase or fumarate reductase, flavoprotein subunit (TIGR01812; HMM-score: 16.4)Energy metabolism Anaerobic succinate dehydrogenase or fumarate reductase, flavoprotein subunit (TIGR01812; HMM-score: 16.4)Energy metabolism Aerobic succinate dehydrogenase or fumarate reductase, flavoprotein subunit (TIGR01812; HMM-score: 16.4)thioredoxin and glutathione reductase (TIGR01438; EC 1.6.4.-; HMM-score: 16)lycopene cyclase family protein (TIGR01790; HMM-score: 15.8)Energy metabolism TCA cycle succinate dehydrogenase, flavoprotein subunit (TIGR01816; HMM-score: 15)Energy metabolism Electron transport flavocytochrome c (TIGR01813; HMM-score: 14.9)Biosynthesis of cofactors, prosthetic groups, and carriers Chlorophyll and bacteriochlorphyll geranylgeranyl reductase (TIGR02028; HMM-score: 12.8)FAD dependent oxidoreductase TIGR03364 (TIGR03364; HMM-score: 12.4)Biosynthesis of cofactors, prosthetic groups, and carriers Menaquinone and ubiquinone ubiquinone biosynthesis hydroxylase, UbiH/UbiF/VisC/COQ6 family (TIGR01988; EC 1.14.13.-; HMM-score: 12.2)Biosynthesis of cofactors, prosthetic groups, and carriers Menaquinone and ubiquinone 2-polyprenyl-6-methoxyphenol 4-hydroxylase (TIGR01984; EC 1.14.13.-; HMM-score: 11.2)Biosynthesis of cofactors, prosthetic groups, and carriers Pyridine nucleotides L-aspartate oxidase (TIGR00551; EC 1.4.3.16; HMM-score: 11.1)Protein synthesis tRNA and rRNA base modification tRNA U-34 5-methylaminomethyl-2-thiouridine biosynthesis protein MnmC, C-terminal domain (TIGR03197; HMM-score: 9.4)Biosynthesis of cofactors, prosthetic groups, and carriers Thiamine glycine oxidase ThiO (TIGR02352; EC 1.4.3.19; HMM-score: 8)
- TheSEED :
- Thioredoxin reductase (EC 1.8.1.9)
and 1 more - PFAM: NADP_Rossmann (CL0063) Pyr_redox_3; Pyridine nucleotide-disulphide oxidoreductase (PF13738; HMM-score: 331.7)and 14 morePyr_redox_2; Pyridine nucleotide-disulphide oxidoreductase (PF07992; HMM-score: 97.5)FAD_oxidored; FAD dependent oxidoreductase (PF12831; HMM-score: 42.3)Pyr_redox; Pyridine nucleotide-disulphide oxidoreductase (PF00070; HMM-score: 38.6)FAD_binding_3; FAD binding domain (PF01494; HMM-score: 35.5)HI0933_like; HI0933-like protein Rossmann domain (PF03486; HMM-score: 34.6)FAD_binding_2; FAD binding domain (PF00890; HMM-score: 33)Thi4; Thi4 family (PF01946; HMM-score: 28.6)NAD_binding_8; NAD(P)-binding Rossmann-like domain (PF13450; HMM-score: 27.3)GIDA; Glucose inhibited division protein A (PF01134; HMM-score: 25.4)DAO; FAD dependent oxidoreductase (PF01266; HMM-score: 25.2)Lycopene_cycl; Lycopene cyclase protein (PF05834; HMM-score: 17.4)FMO-like; Flavin-binding monooxygenase-like (PF00743; HMM-score: 16.9)NAD_binding_7; Putative NAD(P)-binding (PF13241; HMM-score: 13.4)ACT (CL0070) ALS_ss_C; Small subunit of acetolactate synthase (PF10369; HMM-score: 11.7)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 7.5
- Cytoplasmic Membrane Score: 1.15
- Cellwall Score: 0.62
- Extracellular Score: 0.73
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.8773
- Cytoplasmic Membrane Score: 0.1186
- Cell wall & surface Score: 0.0001
- Extracellular Score: 0.0039
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: -1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.014127
- TAT(Tat/SPI): 0.000471
- LIPO(Sec/SPII): 0.009747
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MQKVESIIIGGGPCGLSAAIEQKRKGIDTLIIEKGNVVESIYNYPTHQTFFSSSDKLSIGDVPFIVEESKPRRNQALVYYREVVKHHQLKVNAFEEVLTVKKMNNKFTITTTKDVYECRFLTIATGYYGQHNTLEVEGADLPKVFHYFKEAHPYFDQDVVIIGGKNSAIDAALELEKAGANVTVLYRGGDYSPSIKPWILPNFTALVNHEKIDMEFNANVTQITEDTVTYEVNGESKTIHNDYVFAMIGYHPDYEFLKSVGIQINTNEFGTAPMYNKETYETNIENCYIAGVIAAGNDANTIFIENGKFHGGIIAQSMLAKKQTPLES
⊟Experimental data[edit | edit source]
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: no polycistronic organisation predicted
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
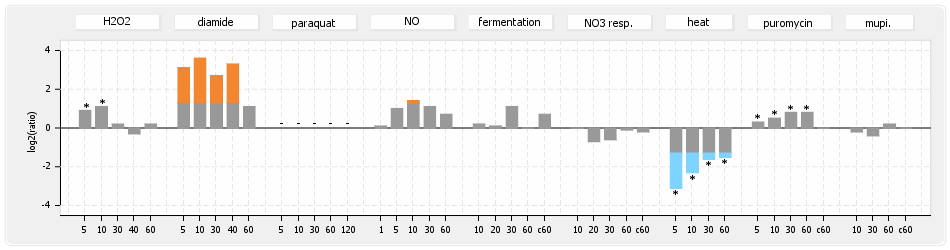
- S.aureus Expression Data Browser: data available for NCTC8325
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: 42.39 h [6]
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Dörte Becher, Kristina Hempel, Susanne Sievers, Daniela Zühlke, Jan Pané-Farré, Andreas Otto, Stephan Fuchs, Dirk Albrecht, Jörg Bernhardt, Susanne Engelmann, Uwe Völker, Jan Maarten van Dijl, Michael Hecker
A proteomic view of an important human pathogen--towards the quantification of the entire Staphylococcus aureus proteome.
PLoS One: 2009, 4(12);e8176
[PubMed:19997597] [WorldCat.org] [DOI] (I e) - ↑ Kristina Hempel, Jan Pané-Farré, Andreas Otto, Susanne Sievers, Michael Hecker, Dörte Becher
Quantitative cell surface proteome profiling for SigB-dependent protein expression in the human pathogen Staphylococcus aureus via biotinylation approach.
J Proteome Res: 2010, 9(3);1579-90
[PubMed:20108986] [WorldCat.org] [DOI] (I p) - ↑ Kristina Hempel, Florian-Alexander Herbst, Martin Moche, Michael Hecker, Dörte Becher
Quantitative proteomic view on secreted, cell surface-associated, and cytoplasmic proteins of the methicillin-resistant human pathogen Staphylococcus aureus under iron-limited conditions.
J Proteome Res: 2011, 10(4);1657-66
[PubMed:21323324] [WorldCat.org] [DOI] (I p) - ↑ Andreas Otto, Jan Maarten van Dijl, Michael Hecker, Dörte Becher
The Staphylococcus aureus proteome.
Int J Med Microbiol: 2014, 304(2);110-20
[PubMed:24439828] [WorldCat.org] [DOI] (I p) - ↑ Daniela Zühlke, Kirsten Dörries, Jörg Bernhardt, Sandra Maaß, Jan Muntel, Volkmar Liebscher, Jan Pané-Farré, Katharina Riedel, Michael Lalk, Uwe Völker, Susanne Engelmann, Dörte Becher, Stephan Fuchs, Michael Hecker
Costs of life - Dynamics of the protein inventory of Staphylococcus aureus during anaerobiosis.
Sci Rep: 2016, 6;28172
[PubMed:27344979] [WorldCat.org] [DOI] (I e) - ↑ Stephan Michalik, Jörg Bernhardt, Andreas Otto, Martin Moche, Dörte Becher, Hanna Meyer, Michael Lalk, Claudia Schurmann, Rabea Schlüter, Holger Kock, Ulf Gerth, Michael Hecker
Life and death of proteins: a case study of glucose-starved Staphylococcus aureus.
Mol Cell Proteomics: 2012, 11(9);558-70
[PubMed:22556279] [WorldCat.org] [DOI] (I p)