Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_02800
- pan locus tag?: SAUPAN006110000
- symbol: SAOUHSC_02800
- pan gene symbol?: sarU
- synonym:
- product: hypothetical protein
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_02800
- symbol: SAOUHSC_02800
- product: hypothetical protein
- replicon: chromosome
- strand: +
- coordinates: 2575879..2576622
- length: 744
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3921454 NCBI
- RefSeq: YP_501259 NCBI
- BioCyc: G1I0R-2638 BioCyc
- MicrobesOnline: 1291230 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721GTGGACTATCAAACTTTCGAAAAGGTCAATAAATTCATAAATGTAGAAGCGTACATATTT
TTTCTCACTCAAGAGTTAAAGCAACAATATAAATTATCATTAAAAGAATTGTTGATATTA
GCATATTTTTATTACAAAAATGAACACAGTATTTCACTAAAAGAAATCATTGGTGACATA
CTTTACAAACAATCTGATGTTGTAAAGAACATTAAGTCACTATCTAAAAAAGGATTTATA
AATAAGTCTAGAAACGAAGCAGATGAACGCCGTATTTTTGTTTCAGTTACTCCAATACAA
CGTAAAAAGATTGCTTGTGTTATTAATGAGTTAGATAAAATAATTAAAGGATTTAATAAG
GAAAGAGACTACATAAAATATCAATGGGCTCCAAAATATAGCAAAGAATTTTTTATACTT
TTTATGAACATTATGTACTCAAAAGATTTTTTAAAATATCGATTTAATTTAACATTTCTT
GATTTATCTATCTTATATGTAATATCATCTCGAAAAAATGAGATACTAAATTTAAAAGAT
TTGTTTGAAAGTATTAGATTTATGTATCCTCAAATTGTTAGGTCAGTTAATAGATTAAAT
AATAAAGGTATGCTAATCAAAGAACGATCCCTTGCAGATGAAAGGATTGTGTTAATCAAA
ATAAATAAAATACAATATAACACTATTAAAAGCATATTCACAGATACTTCCAAGATTCTC
AAACCAAGAAAATTTTTCTTTTAA60
120
180
240
300
360
420
480
540
600
660
720
744
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_02800
- symbol: SAOUHSC_02800
- description: hypothetical protein
- length: 247
- theoretical pI: 10.3367
- theoretical MW: 29792.9
- GRAVY: -0.245344
⊟Function[edit | edit source]
- TIGRFAM: Regulatory functions DNA interactions staphylococcal accessory regulator family (TIGR01889; HMM-score: 210.6)and 1 morehomoprotocatechuate degradation operon regulator, HpaR (TIGR02337; HMM-score: 35.4)
- TheSEED :
- Transcriptional regulator SarU (Staphylococcal accessory regulator U)
- PFAM: HTH (CL0123) Staph_reg_Sar_Rot; Transcriptional regulator SarA/Rot (PF22381; HMM-score: 115.7)and 8 moreHTH_27; Winged helix DNA-binding domain (PF13463; HMM-score: 42.2)MarR_2; MarR family (PF12802; HMM-score: 40)MarR; MarR family (PF01047; HMM-score: 36.4)DnaD_N; DnaD N-terminal domain (PF21984; HMM-score: 19.7)TrmB; Sugar-specific transcriptional regulator TrmB (PF01978; HMM-score: 17.7)F-93_WHD; F-93, winged-helix domain (PF22194; HMM-score: 14.8)WHD_MCM8; MCM8 winged helix domain (PF25051; HMM-score: 13.7)HTH_Tnp_4; Helix-turn-helix of DDE superfamily endonuclease (PF13613; HMM-score: 13.4)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.67
- Cytoplasmic Membrane Score: 0.01
- Cellwall Score: 0.15
- Extracellular Score: 0.17
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.701
- Cytoplasmic Membrane Score: 0.1421
- Cell wall & surface Score: 0.0008
- Extracellular Score: 0.1561
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: -1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.002072
- TAT(Tat/SPI): 0.000099
- LIPO(Sec/SPII): 0.000363
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MDYQTFEKVNKFINVEAYIFFLTQELKQQYKLSLKELLILAYFYYKNEHSISLKEIIGDILYKQSDVVKNIKSLSKKGFINKSRNEADERRIFVSVTPIQRKKIACVINELDKIIKGFNKERDYIKYQWAPKYSKEFFILFMNIMYSKDFLKYRFNLTFLDLSILYVISSRKNEILNLKDLFESIRFMYPQIVRSVNRLNNKGMLIKERSLADERIVLIKINKIQYNTIKSIFTDTSKILKPRKFFF
⊟Experimental data[edit | edit source]
- experimentally validated:
- protein localization:
- quantitative data / protein copy number per cell:
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: no polycistronic organisation predicted
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [1]
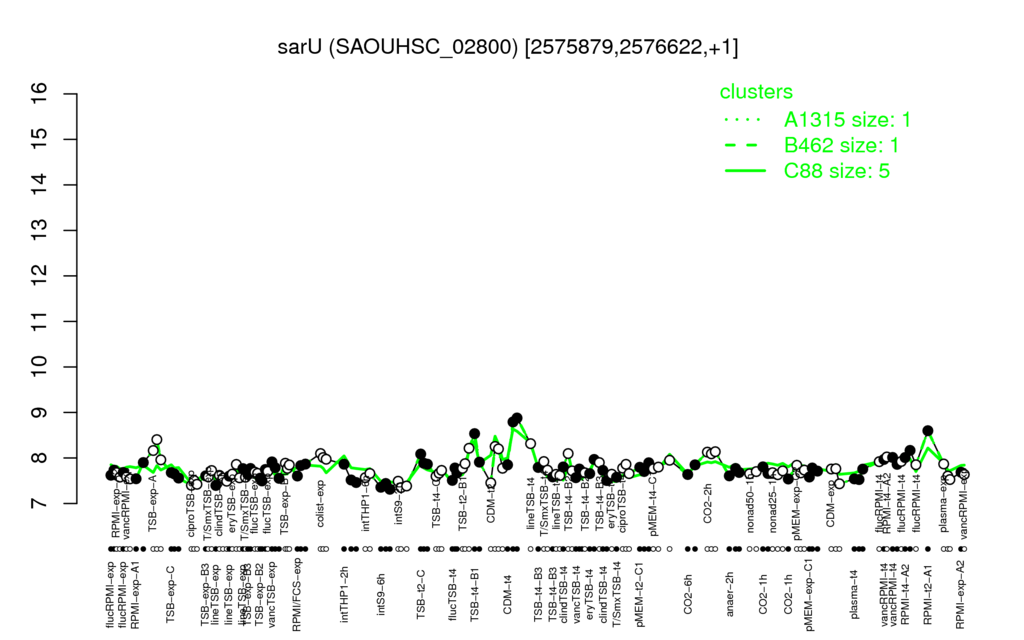 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
⊟Relevant publications[edit | edit source]
Adhar C Manna, Ambrose L Cheung
sarU, a sarA homolog, is repressed by SarT and regulates virulence genes in Staphylococcus aureus.
Infect Immun: 2003, 71(1);343-53
[PubMed:12496184] [WorldCat.org] [DOI] (P p)
