Jump to navigation
Jump to search
NCBI: 10-JUN-2013
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus COL
- locus tag: SACOL1666 [new locus tag: SACOL_RS08500 ]
- pan locus tag?: SAUPAN004207000
- symbol: udk
- pan gene symbol?: udk
- synonym:
- product: uridine kinase
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SACOL1666 [new locus tag: SACOL_RS08500 ]
- symbol: udk
- product: uridine kinase
- replicon: chromosome
- strand: -
- coordinates: 1694389..1695012
- length: 624
- essential: unknown other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3236729 NCBI
- RefSeq: YP_186506 NCBI
- BioCyc: see SACOL_RS08500
- MicrobesOnline: 913115 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601ATGAAAGCTACTACAATCATTGGCATAGCTGGTGGATCTGGCTCAGGAAAAACAACTGTA
ACTAACGAAATTATGAAAAACTTAGAAGGTCATAGTGTCGCTTTACTTGCTCAAGATTAC
TATTATAAAGATCAAAAGCACTTGACTTTCGACGAGCGCCTAGAAACCAATTATGACCAT
CCATTTGCATTCGATAATGATTTATTAATTGAAAATCTTAAAGACTTGAAAAATGGTAAA
GCAGTAGAAGTACCGACATATGATTATGCTAGTCATACAAGAAGTGACATTACCATTGAT
TTTAAACCTAAAGATGTTATTATCGTAGAAGGCATTTTCGCTTTAGAAAATAAGGTATTA
CGTGATATGATGGATGTTAAAATATATGTTGATACAGATGCAGACTTGAGAATATTACGC
CGTTTAACACGAGATACTAAAGAGCGTGGGCGTTCAATGGACTCTGTTATCAATCAATAT
TTAAGTGTTGTTAGACCTATGCATGACCAATTTATTGAACCGACTAAGAAATATGCTGAT
ATAATTATTCCTGAAGGTGGGAGCAATAAAGTTGCAATAGATATTATGACAACAAAAATT
CAGTCTTTAGTTAGCAAGCAATAG60
120
180
240
300
360
420
480
540
600
624
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SACOL1666 [new locus tag: SACOL_RS08500 ]
- symbol: Udk
- description: uridine kinase
- length: 207
- theoretical pI: 6.03587
- theoretical MW: 23504.7
- GRAVY: -0.395652
⊟Function[edit | edit source]
- reaction: EC 2.7.1.48? ExPASyUridine kinase ATP + uridine = ADP + UMP ATP + cytidine = ADP + CMP?
- TIGRFAM: Purines, pyrimidines, nucleosides, and nucleotides Salvage of nucleosides and nucleotides uridine kinase (TIGR00235; EC 2.7.1.48; HMM-score: 294.6)and 23 moreBiosynthesis of cofactors, prosthetic groups, and carriers Pantothenate and coenzyme A pantothenate kinase (TIGR00554; EC 2.7.1.33; HMM-score: 36.2)putative cytidylate kinase (TIGR02173; EC 2.7.4.14; HMM-score: 26.1)Purines, pyrimidines, nucleosides, and nucleotides Nucleotide and nucleoside interconversions cytidylate kinase (TIGR00017; EC 2.7.4.14; HMM-score: 20.6)Biosynthesis of cofactors, prosthetic groups, and carriers Molybdopterin molybdopterin-guanine dinucleotide biosynthesis protein B (TIGR00176; HMM-score: 19.9)Biosynthesis of cofactors, prosthetic groups, and carriers Pantothenate and coenzyme A dephospho-CoA kinase (TIGR00152; EC 2.7.1.24; HMM-score: 18.9)Cellular processes Toxin production and resistance putative bacteriocin export ABC transporter, lactococcin 972 group (TIGR03608; HMM-score: 18.5)Transport and binding proteins Unknown substrate putative bacteriocin export ABC transporter, lactococcin 972 group (TIGR03608; HMM-score: 18.5)Central intermediary metabolism Nitrogen metabolism urease accessory protein UreG (TIGR00101; HMM-score: 17.3)Purines, pyrimidines, nucleosides, and nucleotides Nucleotide and nucleoside interconversions dTMP kinase (TIGR00041; EC 2.7.4.9; HMM-score: 16.3)Central intermediary metabolism Sulfur metabolism adenylyl-sulfate kinase (TIGR00455; EC 2.7.1.25; HMM-score: 15.5)Transport and binding proteins Cations and iron carrying compounds nickel import ATP-binding protein NikD (TIGR02770; HMM-score: 14.6)P-type DNA transfer ATPase VirB11 (TIGR02788; HMM-score: 14.5)Cell envelope Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides cellulose synthase operon protein YhjQ (TIGR03371; HMM-score: 13.7)Biosynthesis of cofactors, prosthetic groups, and carriers Heme, porphyrin, and cobalamin cobalamin biosynthesis protein CobW (TIGR02475; HMM-score: 13.2)DNA metabolism DNA replication, recombination, and repair orc1/cdc6 family replication initiation protein (TIGR02928; HMM-score: 12.9)Transport and binding proteins Amino acids, peptides and amines LAO/AO transport system ATPase (TIGR00750; EC 2.7.-.-; HMM-score: 12.5)Regulatory functions Protein interactions LAO/AO transport system ATPase (TIGR00750; EC 2.7.-.-; HMM-score: 12.5)Cellular processes Conjugation P-type conjugative transfer ATPase TrbB (TIGR02782; HMM-score: 12.3)Central intermediary metabolism Phosphorus compounds phosphonate C-P lyase system protein PhnK (TIGR02323; HMM-score: 11.1)Protein fate Protein and peptide secretion and trafficking type I secretion system ATPase (TIGR01846; HMM-score: 10.8)thiol reductant ABC exporter, CydD subunit (TIGR02857; HMM-score: 10.5)thiol reductant ABC exporter, CydC subunit (TIGR02868; HMM-score: 10)Transport and binding proteins Other pigment precursor permease (TIGR00955; HMM-score: 8.7)
- TheSEED :
- Uridine kinase (EC 2.7.1.48)
- PFAM: P-loop_NTPase (CL0023) PRK; Phosphoribulokinase / Uridine kinase family (PF00485; HMM-score: 163.1)and 26 moreAAA_18; AAA domain (PF13238; HMM-score: 33.7)AAA_17; AAA domain (PF13207; HMM-score: 28.3)dNK; Deoxynucleoside kinase (PF01712; HMM-score: 24.8)ABC_tran; ABC transporter (PF00005; HMM-score: 21.2)CoaE; Dephospho-CoA kinase (PF01121; HMM-score: 20.6)cobW; CobW/HypB/UreG, nucleotide-binding domain (PF02492; HMM-score: 20.1)Cytidylate_kin; Cytidylate kinase (PF02224; HMM-score: 20)NTPase_1; NTPase (PF03266; HMM-score: 18.4)AAA_19; AAA domain (PF13245; HMM-score: 17.5)AAA_28; AAA domain (PF13521; HMM-score: 16.5)RNA_helicase; RNA helicase (PF00910; HMM-score: 16.4)AAA_16; AAA ATPase domain (PF13191; HMM-score: 16.4)MobB; Molybdopterin guanine dinucleotide synthesis protein B (PF03205; HMM-score: 15)MeaB; Methylmalonyl Co-A mutase-associated GTPase MeaB (PF03308; HMM-score: 14.8)AAA_33; AAA domain (PF13671; HMM-score: 14.8)AAA_22; AAA domain (PF13401; HMM-score: 14.6)APS_kinase; Adenylylsulphate kinase (PF01583; HMM-score: 14.1)Zeta_toxin; Zeta toxin (PF06414; HMM-score: 14)CbiA; CobQ/CobB/MinD/ParA nucleotide binding domain (PF01656; HMM-score: 13.7)Thymidylate_kin; Thymidylate kinase (PF02223; HMM-score: 13.6)NPHP3_N; Nephrocystin 3, N-terminal (PF24883; HMM-score: 13.6)ResIII; Type III restriction enzyme, res subunit (PF04851; HMM-score: 13.5)AAA_14; AAA domain (PF13173; HMM-score: 13.2)NB-ARC; NB-ARC domain (PF00931; HMM-score: 12.8)TrwB_AAD_bind; Type IV secretion-system coupling protein DNA-binding domain (PF10412; HMM-score: 12)AAA_30; AAA domain (PF13604; HMM-score: 11.6)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.67
- Cytoplasmic Membrane Score: 0.01
- Cellwall Score: 0.15
- Extracellular Score: 0.17
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.9811
- Cytoplasmic Membrane Score: 0.0014
- Cell wall & surface Score: 0
- Extracellular Score: 0.0174
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: -1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.044006
- TAT(Tat/SPI): 0.0012
- LIPO(Sec/SPII): 0.033459
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MKATTIIGIAGGSGSGKTTVTNEIMKNLEGHSVALLAQDYYYKDQKHLTFDERLETNYDHPFAFDNDLLIENLKDLKNGKAVEVPTYDYASHTRSDITIDFKPKDVIIVEGIFALENKVLRDMMDVKIYVDTDADLRILRRLTRDTKERGRSMDSVINQYLSVVRPMHDQFIEPTKKYADIIIPEGGSNKVAIDIMTTKIQSLVSKQ
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas
- protein localization: Cytoplasmic [1] [2] [3]
- quantitative data / protein copy number per cell: 408 [4]
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: data available for NCTC8325
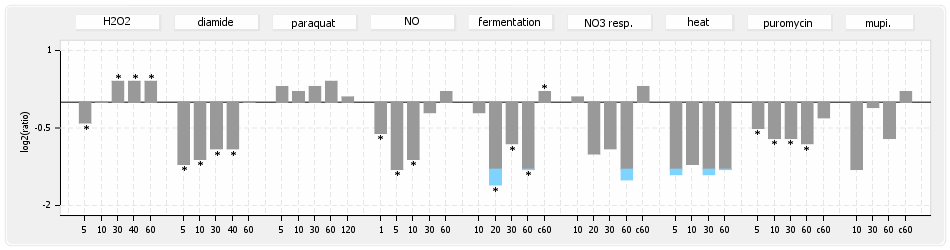
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: 25.25 h [5]
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Dörte Becher, Kristina Hempel, Susanne Sievers, Daniela Zühlke, Jan Pané-Farré, Andreas Otto, Stephan Fuchs, Dirk Albrecht, Jörg Bernhardt, Susanne Engelmann, Uwe Völker, Jan Maarten van Dijl, Michael Hecker
A proteomic view of an important human pathogen--towards the quantification of the entire Staphylococcus aureus proteome.
PLoS One: 2009, 4(12);e8176
[PubMed:19997597] [WorldCat.org] [DOI] (I e) - ↑ Kristina Hempel, Florian-Alexander Herbst, Martin Moche, Michael Hecker, Dörte Becher
Quantitative proteomic view on secreted, cell surface-associated, and cytoplasmic proteins of the methicillin-resistant human pathogen Staphylococcus aureus under iron-limited conditions.
J Proteome Res: 2011, 10(4);1657-66
[PubMed:21323324] [WorldCat.org] [DOI] (I p) - ↑ Andreas Otto, Jan Maarten van Dijl, Michael Hecker, Dörte Becher
The Staphylococcus aureus proteome.
Int J Med Microbiol: 2014, 304(2);110-20
[PubMed:24439828] [WorldCat.org] [DOI] (I p) - ↑ Daniela Zühlke, Kirsten Dörries, Jörg Bernhardt, Sandra Maaß, Jan Muntel, Volkmar Liebscher, Jan Pané-Farré, Katharina Riedel, Michael Lalk, Uwe Völker, Susanne Engelmann, Dörte Becher, Stephan Fuchs, Michael Hecker
Costs of life - Dynamics of the protein inventory of Staphylococcus aureus during anaerobiosis.
Sci Rep: 2016, 6;28172
[PubMed:27344979] [WorldCat.org] [DOI] (I e) - ↑ Stephan Michalik, Jörg Bernhardt, Andreas Otto, Martin Moche, Dörte Becher, Hanna Meyer, Michael Lalk, Claudia Schurmann, Rabea Schlüter, Holger Kock, Ulf Gerth, Michael Hecker
Life and death of proteins: a case study of glucose-starved Staphylococcus aureus.
Mol Cell Proteomics: 2012, 11(9);558-70
[PubMed:22556279] [WorldCat.org] [DOI] (I p)