Jump to navigation
Jump to search
NCBI: 10-JUN-2013
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus COL
- locus tag: SACOL2045 [new locus tag: SACOL_RS10680 ]
- pan locus tag?: SAUPAN005306000
- symbol: ilvC
- pan gene symbol?: ilvC
- synonym:
- product: ketol-acid reductoisomerase
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SACOL2045 [new locus tag: SACOL_RS10680 ]
- symbol: ilvC
- product: ketol-acid reductoisomerase
- replicon: chromosome
- strand: +
- coordinates: 2104945..2105949
- length: 1005
- essential: unknown other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3238699 NCBI
- RefSeq: YP_186862 NCBI
- BioCyc: see SACOL_RS10680
- MicrobesOnline: 913518 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961ATGACAACAGTTTATTATGATCAAGATGTAAAAACGGACGCTTTACAAGGCAAAAAAATT
GCAGTAGTAGGTTATGGATCACAAGGTCACGCGCATGCACAAAACTTAAAAGACAATGGA
TATGATGTAGTCATCGGCATTCGCCCAGGTCGTTCTTTTGACAAAGCTAAAGAAGATGGA
TTTGATGTGTTCCCTGTTGCAGAAGCAGTTAAGCAAGCTGATGTAATTATGGTGCTATTA
CCTGATGAAATTCAAGGTGATGTATACAAAAACGAAATTGAACCAAATTTAGAAAAACAT
AATGCGCTTGCATTTGCTCATGGCTTTAACATTCATTTTGGTGTTATTCAACCACCAGCT
GATGTTGATGTATTTTTAGTAGCTCCTAAAGGACCGGGTCATTTAGTTAGACGTACATTT
GTTGAAGGTTCTGCTGTACCATCACTATTTGGTATTCAACAAGGCGCTTCAGGTCAAGCA
CGTAATATTGCTTTAAGTTATGCAAAAGGTATTGGTGCAACTCGTGCAGGTGTTATTGAA
ACAACATTTAAAGAAGAAACTGAGACAGATTTATTTGGTGAACAAGCAGTACTTTGCGGT
GGTGTATCGAAATTAATTCAAAGTGGCTTTGAAACATTAGTAGAAGCGGGTTATCAACCA
GAATTAGCTTATTTTGAAGTATTACATGAAATGAAATTAATCGTTGATTTGATGTATGAA
GGCGGTATGGAAAATGTACGTTACTCAATTTCAAATACTGCTGAATTTGGTGACTATGTT
TCAGGACCACGTGTTATCACACCAGATGTTAAAGAAAATATGAAAGCTGTATTAACTGAT
ATCCAAAATGGTAACTTCAGTAATCGCTTTATCGAAGACAATAAAAATGGATTCAAAGAA
TTTTATAAATTACGCGAAGAACAACATGGTCATCAAATTGAAAAAGTTGGTCGTGAATTA
CGCGAAATGATGCCTTTTATTAAATCTAAAAGCATTGAAAAATAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1005
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SACOL2045 [new locus tag: SACOL_RS10680 ]
- symbol: IlvC
- description: ketol-acid reductoisomerase
- length: 334
- theoretical pI: 5.04028
- theoretical MW: 36955.6
- GRAVY: -0.29491
⊟Function[edit | edit source]
- reaction: EC 1.1.1.86? ExPASyKetol-acid reductoisomerase (NADP+) (2R)-2,3-dihydroxy-3-methylbutanoate + NADP+ = (2S)-2-hydroxy-2-methyl-3-oxobutanoate + NADPH (2R,3R)-2,3-dihydroxy-3-methylpentanoate + NADP+ = (S)-2-hydroxy-2-ethyl-3-oxobutanoate + NADPH
- TIGRFAM: Amino acid biosynthesis Pyruvate family ketol-acid reductoisomerase (TIGR00465; EC 1.1.1.86; HMM-score: 436.7)and 7 moreEnergy metabolism Amino acids and amines adenosylhomocysteinase (TIGR00936; EC 3.3.1.1; HMM-score: 31.9)2-hydroxy-3-oxopropionate reductase (TIGR01505; EC 1.1.1.60; HMM-score: 17.5)Cellular processes Sporulation and germination dipicolinic acid synthetase, A subunit (TIGR02853; HMM-score: 15.6)Biosynthesis of cofactors, prosthetic groups, and carriers Pantothenate and coenzyme A 2-dehydropantoate 2-reductase (TIGR00745; EC 1.1.1.-; HMM-score: 15.3)Energy metabolism Amino acids and amines 3-hydroxyisobutyrate dehydrogenase (TIGR01692; EC 1.1.1.31; HMM-score: 14.5)Energy metabolism Pentose phosphate pathway 6-phosphogluconate dehydrogenase (decarboxylating) (TIGR00872; EC 1.1.1.44; HMM-score: 12.7)glutamate synthase, NADH/NADPH, small subunit (TIGR01317; EC 1.4.1.-; HMM-score: 12.4)
- TheSEED :
- Ketol-acid reductoisomerase (EC 1.1.1.86)
Amino Acids and Derivatives Branched-chain amino acids Branched-Chain Amino Acid Biosynthesis Ketol-acid reductoisomerase (EC 1.1.1.86)and 1 more - PFAM: NADP_Rossmann (CL0063) KARI_N; Acetohydroxy acid isomeroreductase, NADPH-binding domain (PF07991; HMM-score: 252)6PGD_C (CL0106) KARI_C; Ketol-acid reductoisomerase, C-terminal domain (PF01450; HMM-score: 217.3)and 11 moreNADP_Rossmann (CL0063) 2-Hacid_dh_C; D-isomer specific 2-hydroxyacid dehydrogenase, NAD binding domain (PF02826; HMM-score: 32.1)F420_oxidored; NADP oxidoreductase coenzyme F420-dependent (PF03807; HMM-score: 29.3)NAD_binding_2; NAD binding domain of 6-phosphogluconate dehydrogenase (PF03446; HMM-score: 27.9)AdoHcyase_NAD; S-adenosyl-L-homocysteine hydrolase, NAD binding domain (PF00670; HMM-score: 27.6)NAD_Gly3P_dh_N; NAD-dependent glycerol-3-phosphate dehydrogenase N-terminus (PF01210; HMM-score: 20)CoA_binding; CoA binding domain (PF02629; HMM-score: 19)ApbA; Ketopantoate reductase PanE/ApbA (PF02558; HMM-score: 15.7)CoA_binding_2; CoA binding domain (PF13380; HMM-score: 14.2)TrkA_N; TrkA-N domain (PF02254; HMM-score: 13.3)Shikimate_DH; Shikimate / quinate 5-dehydrogenase (PF01488; HMM-score: 12.8)Chelatase (CL0043) Oxidored_nitro; Nitrogenase component 1 type Oxidoreductase (PF00148; HMM-score: 11.8)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors: Mg2+
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.97
- Cytoplasmic Membrane Score: 0
- Cellwall Score: 0.01
- Extracellular Score: 0.02
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.8897
- Cytoplasmic Membrane Score: 0.0078
- Cell wall & surface Score: 0.0006
- Extracellular Score: 0.1019
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.059023
- TAT(Tat/SPI): 0.00266
- LIPO(Sec/SPII): 0.005356
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MTTVYYDQDVKTDALQGKKIAVVGYGSQGHAHAQNLKDNGYDVVIGIRPGRSFDKAKEDGFDVFPVAEAVKQADVIMVLLPDEIQGDVYKNEIEPNLEKHNALAFAHGFNIHFGVIQPPADVDVFLVAPKGPGHLVRRTFVEGSAVPSLFGIQQGASGQARNIALSYAKGIGATRAGVIETTFKEETETDLFGEQAVLCGGVSKLIQSGFETLVEAGYQPELAYFEVLHEMKLIVDLMYEGGMENVRYSISNTAEFGDYVSGPRVITPDVKENMKAVLTDIQNGNFSNRFIEDNKNGFKEFYKLREEQHGHQIEKVGRELREMMPFIKSKSIEK
⊟Experimental data[edit | edit source]
⊟Expression & Regulation[edit | edit source]
⊟Regulation[edit | edit source]
- regulator: CodY (repression) regulon
CodY (TF) important in Amino acid metabolism; RegPrecise transcription unit transferred from N315 data RegPrecise
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: data available for NCTC8325
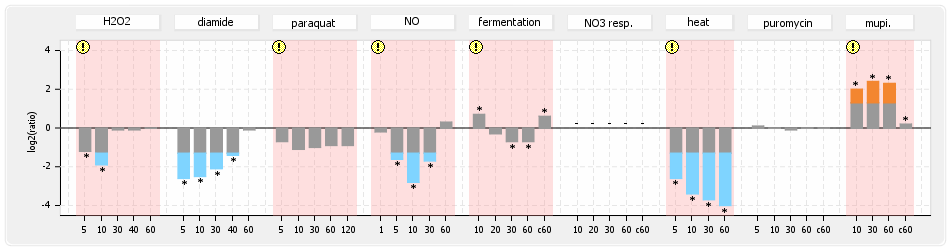
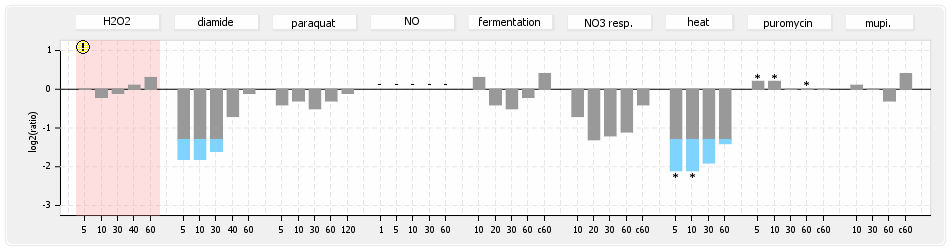
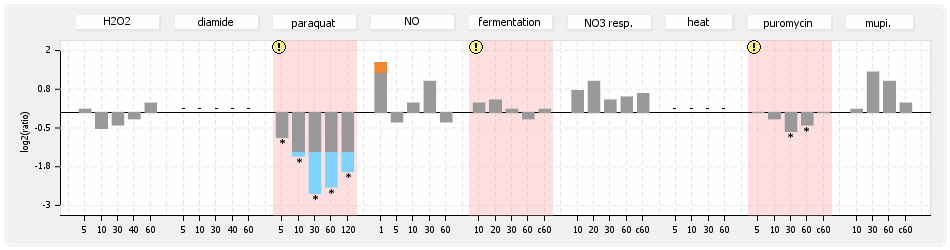
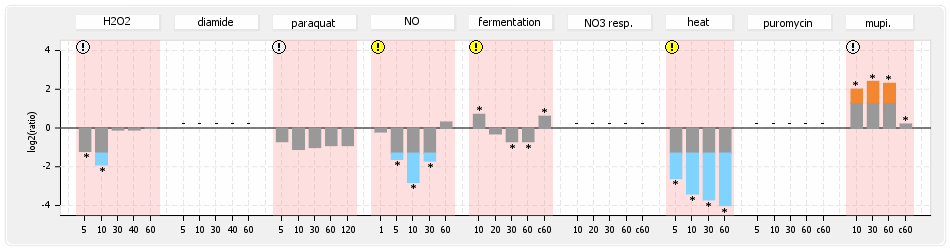
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: 45.11 h [8]
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Dörte Becher, Kristina Hempel, Susanne Sievers, Daniela Zühlke, Jan Pané-Farré, Andreas Otto, Stephan Fuchs, Dirk Albrecht, Jörg Bernhardt, Susanne Engelmann, Uwe Völker, Jan Maarten van Dijl, Michael Hecker
A proteomic view of an important human pathogen--towards the quantification of the entire Staphylococcus aureus proteome.
PLoS One: 2009, 4(12);e8176
[PubMed:19997597] [WorldCat.org] [DOI] (I e) - ↑ Kristina Hempel, Jan Pané-Farré, Andreas Otto, Susanne Sievers, Michael Hecker, Dörte Becher
Quantitative cell surface proteome profiling for SigB-dependent protein expression in the human pathogen Staphylococcus aureus via biotinylation approach.
J Proteome Res: 2010, 9(3);1579-90
[PubMed:20108986] [WorldCat.org] [DOI] (I p) - ↑ Annette Dreisbach, Kristina Hempel, Girbe Buist, Michael Hecker, Dörte Becher, Jan Maarten van Dijl
Profiling the surfacome of Staphylococcus aureus.
Proteomics: 2010, 10(17);3082-96
[PubMed:20662103] [WorldCat.org] [DOI] (I p) - ↑ Kristina Hempel, Florian-Alexander Herbst, Martin Moche, Michael Hecker, Dörte Becher
Quantitative proteomic view on secreted, cell surface-associated, and cytoplasmic proteins of the methicillin-resistant human pathogen Staphylococcus aureus under iron-limited conditions.
J Proteome Res: 2011, 10(4);1657-66
[PubMed:21323324] [WorldCat.org] [DOI] (I p) - ↑ Andreas Otto, Jan Maarten van Dijl, Michael Hecker, Dörte Becher
The Staphylococcus aureus proteome.
Int J Med Microbiol: 2014, 304(2);110-20
[PubMed:24439828] [WorldCat.org] [DOI] (I p) - ↑ Daniela Zühlke, Kirsten Dörries, Jörg Bernhardt, Sandra Maaß, Jan Muntel, Volkmar Liebscher, Jan Pané-Farré, Katharina Riedel, Michael Lalk, Uwe Völker, Susanne Engelmann, Dörte Becher, Stephan Fuchs, Michael Hecker
Costs of life - Dynamics of the protein inventory of Staphylococcus aureus during anaerobiosis.
Sci Rep: 2016, 6;28172
[PubMed:27344979] [WorldCat.org] [DOI] (I e) - ↑ Charlotte D Majerczyk, Paul M Dunman, Thanh T Luong, Chia Y Lee, Marat R Sadykov, Greg A Somerville, Kip Bodi, Abraham L Sonenshein
Direct targets of CodY in Staphylococcus aureus.
J Bacteriol: 2010, 192(11);2861-77
[PubMed:20363936] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Jörg Bernhardt, Andreas Otto, Martin Moche, Dörte Becher, Hanna Meyer, Michael Lalk, Claudia Schurmann, Rabea Schlüter, Holger Kock, Ulf Gerth, Michael Hecker
Life and death of proteins: a case study of glucose-starved Staphylococcus aureus.
Mol Cell Proteomics: 2012, 11(9);558-70
[PubMed:22556279] [WorldCat.org] [DOI] (I p)