Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_00150
- pan locus tag?: SAUPAN001025000
- symbol: SAOUHSC_00150
- pan gene symbol?: argD
- synonym: rocD
- product: ornithine aminotransferase
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_00150
- symbol: SAOUHSC_00150
- product: ornithine aminotransferase
- replicon: chromosome
- strand: -
- coordinates: 161057..162241
- length: 1185
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3919858 NCBI
- RefSeq: YP_498749 NCBI
- BioCyc: G1I0R-140 BioCyc
- MicrobesOnline: 1288643 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141ATGAATTCAATCATTGAATTAACTGATTATTATAGCTCTAATAATTATGCACCACTTAAG
CTTGTCATTTCTAAAGGTAAAGGTGTCAAAGTTTGGGATACTGATGGCAAACAATATATA
GATTGCATTTCGGGTTTTTCAGTTGCAAACCAAGGCCATTGTCATCCAACAATTGTTAAA
GCGATGACAGAACAAGCTTCAAAGTTGTCTATCATTTCACGTGTCCTTTATAGTGACAAT
CTCGGGAAATGGGAAGAAAAAATTTGTCATCTTGCTAAGAAAGACAAAGTACTCCCCCTT
AACTCTGGTACTGAAGCTGTTGAAGCAGCCATTAAAATTGCTAGAAAATGGGGCTCTGAA
GTTAAAGGCATTACTGACGGACAAGTTGAAATCATCGCTATGAATAACAATTTTCACGGT
CGTACACTTGGCTCATTATCACTATCTAACCACGACGCATATAAAGCAGGATTTCACCCC
CTACTTCAAGGCACTACAACAGTAGATTTTGGAGACATTGAACAATTAACACAAGCTATT
TCACCGAATACAGCAGCAATTATTTTGGAACCAATTCAAGGTGAAGGTGGCGTTAATATA
CCACCGAAAGGATATATTCAAGCTGTGCGTCAACTATGTGATAAACATCAAATATTATTG
ATTGCAGATGAAATTCAAGTTGGTCTTGGTAGAACTGGGAAATGGTTTGCTATGGAATGG
GAGCAAGTCGTTCCAGACATTTATATTTTAGGTAAGGCATTGGGTGGCGGCTTATACCCT
GTATCTGCTGTACTTGCAAATAATGATGTCATGCGTGTTCTAACACCAGGTACACATGGT
TCAACATTTGGTGGTAACCCTTTAGCCATTGCAATATCGACGGCAGCGCTTGATGTACTT
AAAGATGAACAACTGGTTGAACGATCAGAACGCTTAGGTTCATTTTTATTAAAAGCGTTG
CTACAACTTAAACATCCTAGTATTAAAGAAATTAGAGGTCGTGGTTTATTTATAGGCATA
GAGCTTAACACAGATGCTGCACCTTTTGTGGATCAACTGATTCAACGTGGAATCTTATGC
AAAGACACGCATCGTACTATCATTCGATTGTCTCCACCTCTAGTCATTGATAAAGAGGAA
ATCCATCAAATTGTTGCAGCTTTTCAAGACGTTTTTAAAAATTAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1185
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_00150
- symbol: SAOUHSC_00150
- description: ornithine aminotransferase
- length: 394
- theoretical pI: 7.10134
- theoretical MW: 43048.3
- GRAVY: -0.0327411
⊟Function[edit | edit source]
- reaction: EC 2.6.1.13? ExPASyOrnithine aminotransferase L-ornithine + a 2-oxo acid = L-glutamate 5-semialdehyde + an L-amino acid
- TIGRFAM: ornithine--oxo-acid transaminase (TIGR01885; EC 2.6.1.13; HMM-score: 486.8)transaminase, acetylornithine/succinylornithine family (TIGR00707; HMM-score: 424.9)and 13 moreEnergy metabolism Amino acids and amines succinylornithine transaminase family (TIGR03246; EC 2.6.1.81; HMM-score: 328)Central intermediary metabolism Polyamine biosynthesis putrescine aminotransferase (TIGR03372; EC 2.6.1.82; HMM-score: 271.3)Biosynthesis of cofactors, prosthetic groups, and carriers Biotin adenosylmethionine-8-amino-7-oxononanoate transaminase (TIGR00508; EC 2.6.1.62; HMM-score: 262.1)Central intermediary metabolism Other 4-aminobutyrate transaminase (TIGR00700; EC 2.6.1.19; HMM-score: 247.9)Central intermediary metabolism Other 2,4-diaminobutyrate 4-transaminase (TIGR00709; EC 2.6.1.-; HMM-score: 216.3)Cellular processes Adaptations to atypical conditions diaminobutyrate--2-oxoglutarate aminotransferase (TIGR02407; EC 2.6.1.76; HMM-score: 198.2)L-lysine 6-transaminase (TIGR03251; EC 2.6.1.36; HMM-score: 160.3)Biosynthesis of cofactors, prosthetic groups, and carriers Heme, porphyrin, and cobalamin glutamate-1-semialdehyde-2,1-aminomutase (TIGR00713; EC 5.4.3.8; HMM-score: 145.5)Central intermediary metabolism Other 4-aminobutyrate aminotransferase (TIGR00699; EC 2.6.1.19; HMM-score: 109.8)Biosynthesis of cofactors, prosthetic groups, and carriers Heme, porphyrin, and cobalamin threonine-phosphate decarboxylase (TIGR01140; EC 4.1.1.81; HMM-score: 14.4)tyrosine/nicotianamine family aminotransferase (TIGR01265; HMM-score: 14.1)Energy metabolism Amino acids and amines tyrosine aminotransferase (TIGR01264; EC 2.6.1.5; HMM-score: 13.4)Amino acid biosynthesis Histidine family histidinol-phosphate transaminase (TIGR01141; EC 2.6.1.9; HMM-score: 12.3)
- TheSEED :
- Acetylornithine aminotransferase (EC 2.6.1.11)
- PFAM: PLP_aminotran (CL0061) Aminotran_3; Aminotransferase class-III (PF00202; HMM-score: 442.6)and 7 moreDegT_DnrJ_EryC1; DegT/DnrJ/EryC1/StrS aminotransferase family (PF01041; HMM-score: 22.6)Beta_elim_lyase; Beta-eliminating lyase (PF01212; HMM-score: 22.1)Aminotran_1_2; Aminotransferase class I and II (PF00155; HMM-score: 20.4)KYNU_C; Kynureninase C-terminal domain (PF22580; HMM-score: 13.8)Cys_Met_Meta_PP; Cys/Met metabolism PLP-dependent enzyme (PF01053; HMM-score: 13.4)HTH (CL0123) MRB1590_C; MRB1590 C-terminal domain (PF21117; HMM-score: 13.4)PLP_aminotran (CL0061) Aminotran_5; Aminotransferase class-V (PF00266; HMM-score: 12.2)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors: pyridoxal 5'-phosphate
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.97
- Cytoplasmic Membrane Score: 0
- Cellwall Score: 0.01
- Extracellular Score: 0.02
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.9765
- Cytoplasmic Membrane Score: 0.0072
- Cell wall & surface Score: 0.0006
- Extracellular Score: 0.0157
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.132451
- TAT(Tat/SPI): 0.000497
- LIPO(Sec/SPII): 0.001728
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MNSIIELTDYYSSNNYAPLKLVISKGKGVKVWDTDGKQYIDCISGFSVANQGHCHPTIVKAMTEQASKLSIISRVLYSDNLGKWEEKICHLAKKDKVLPLNSGTEAVEAAIKIARKWGSEVKGITDGQVEIIAMNNNFHGRTLGSLSLSNHDAYKAGFHPLLQGTTTVDFGDIEQLTQAISPNTAAIILEPIQGEGGVNIPPKGYIQAVRQLCDKHQILLIADEIQVGLGRTGKWFAMEWEQVVPDIYILGKALGGGLYPVSAVLANNDVMRVLTPGTHGSTFGGNPLAIAISTAALDVLKDEQLVERSERLGSFLLKALLQLKHPSIKEIRGRGLFIGIELNTDAAPFVDQLIQRGILCKDTHRTIIRLSPPLVIDKEEIHQIVAAFQDVFKN
⊟Experimental data[edit | edit source]
- experimentally validated:
- protein localization:
- quantitative data / protein copy number per cell:
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
⊟Regulation[edit | edit source]
- regulator: ArgR*/AhrC (repression) regulon
ArgR*/AhrC (TF) important in Arginine biosynthesis, Arginine degradation; RegPrecise
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [1]
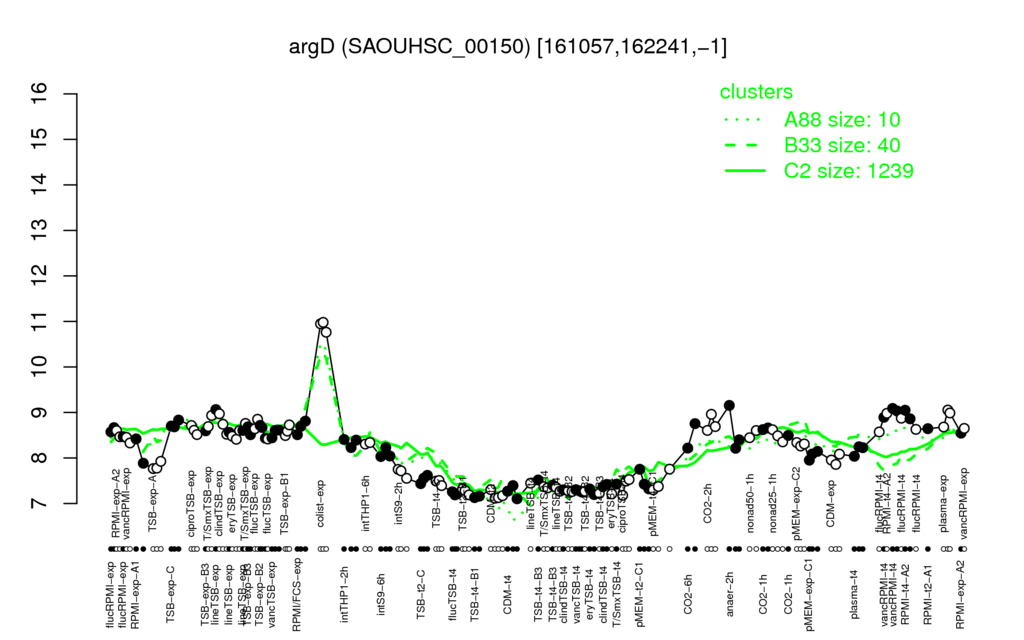 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
