Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_00219
- pan locus tag?: SAUPAN001122000
- symbol: SAOUHSC_00219
- pan gene symbol?: —
- synonym:
- product: hypothetical protein
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_00219
- symbol: SAOUHSC_00219
- product: hypothetical protein
- replicon: chromosome
- strand: +
- coordinates: 238356..239399
- length: 1044
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3920295 NCBI
- RefSeq: YP_498814 NCBI
- BioCyc: G1I0R-204 BioCyc
- MicrobesOnline: 1288708 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021GTGAAAGCATTGAAATTATATGGCGTGGAAGATTTACGGTATGAGGATAATGAAAAGCCA
GTCATTGAAAGTGCGAATGACGTTATTATTAAAGTACGAGCGACTGGCATATGTGGTTCA
GACACGTCACGATACAAAAAAATGGGGCCATACATTAAAGGTATGCCATTTGGTCATGAA
TTTTCAGGTGTAGTAGATGCCATTGGAAGTGATGTTACGCATGTTAATGTGGGCGACAAA
GTGACAGGTTGCCCAGCAATACCTTGTTATCAATGCGAGTATTGTTTAAAAGGTGAATAT
GCACGATGTGAAAAGTTATTCGTCATTGGCTCATATGAACCTGGATCGTTCGCGGAATAT
GTCAAATTGCCAGCGCAAAATGTTTTAAAGGTTCCAGACAATGTTGATTACATTGAAGCA
GCAATGGTTGAGCCATCAGCCGTTGTTGCGCATGGGTTTTATAAATCGAATATACAACCT
GGTATGACTGTTGCAGTAATGGGGTGTGGCAGTATAGGTTTGTTAGCTATTCAATGGGCA
CGAATATTTGGTGCTGCACATATCATCGCTATAGATATAGATGCGCATAAACTAGATATT
GCAACATCATTGGGCGCACATCAAACAATCAATTCAAAAGAAGAAAATCTTGAGAAATTC
ATCGAAAATCATTACGCCAATCAAATCGATTTAGCTATAGAATCATCAGGTGCTAAAGTT
ACGATTGGTCAAATATTGACGCTACCTAAAAAAGGTGGCGAGGTGGTATTACTCGGAATA
CCATATGATGATATTGAGATTGATCGCGTTCATTTTGAAAAAATTCTGCGTAACGAGTTG
ACAGTATGTGGCTCTTGGAACTGTTTGTCCAGTAATTTTCCGGGCAAAGAGTGGACGGCA
ACCTTACATTATATGAAGACGAAAGATATTAATGTAAAGCCTATTATTTCTCATTTTTTA
CCGTTAGAAAAAGGCCCGGAGACATTTGATAAATTAGTTAACAAGAAAGAACGATTTGAT
AAAGTCATGTTTACGATTTATTAG60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1044
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_00219
- symbol: SAOUHSC_00219
- description: hypothetical protein
- length: 347
- theoretical pI: 6.1837
- theoretical MW: 38401.1
- GRAVY: -0.0694524
⊟Function[edit | edit source]
- TIGRFAM: Energy metabolism Amino acids and amines L-threonine 3-dehydrogenase (TIGR00692; EC 1.1.1.103; HMM-score: 198.1)and 17 moreCellular processes Detoxification S-(hydroxymethyl)mycothiol dehydrogenase (TIGR03451; EC 1.1.1.306; HMM-score: 121)putative phosphonate catabolism associated alcohol dehydrogenase (TIGR03366; HMM-score: 117.5)Unknown function Enzymes of unknown specificity NDMA-dependent alcohol dehydrogenase, Rxyl_3153 family (TIGR03989; EC 1.1.99.36; HMM-score: 116.9)6-hydroxycyclohex-1-ene-1-carbonyl-CoA dehydrogenase (TIGR03201; EC 1.1.1.-; HMM-score: 108.8)Cellular processes Detoxification S-(hydroxymethyl)glutathione dehydrogenase/class III alcohol dehydrogenase (TIGR02818; EC 1.1.1.1,1.1.1.284; HMM-score: 102.6)Energy metabolism Fermentation S-(hydroxymethyl)glutathione dehydrogenase/class III alcohol dehydrogenase (TIGR02818; EC 1.1.1.1,1.1.1.284; HMM-score: 102.6)Unknown function Enzymes of unknown specificity putative NAD(P)H quinone oxidoreductase, PIG3 family (TIGR02824; HMM-score: 97.6)Central intermediary metabolism One-carbon metabolism formaldehyde dehydrogenase, glutathione-independent (TIGR02819; EC 1.2.1.46; HMM-score: 77.3)Energy metabolism Fermentation zinc-binding alcohol dehydrogenase family protein (TIGR02822; EC 1.-.-.-; HMM-score: 56.3)crotonyl-CoA carboxylase/reductase (TIGR01751; EC 1.3.1.85; HMM-score: 50.2)Energy metabolism Fermentation zinc-binding alcohol dehydrogenase family protein (TIGR02817; HMM-score: 42)Unknown function Enzymes of unknown specificity putative quinone oxidoreductase, YhdH/YhfP family (TIGR02823; HMM-score: 33.3)Biosynthesis of cofactors, prosthetic groups, and carriers Chlorophyll and bacteriochlorphyll chlorophyll synthesis pathway protein BchC (TIGR01202; HMM-score: 32.8)Protein synthesis Ribosomal proteins: synthesis and modification ribosomal protein L11 methyltransferase (TIGR00406; EC 2.1.1.-; HMM-score: 20.1)Biosynthesis of cofactors, prosthetic groups, and carriers Heme, porphyrin, and cobalamin precorrin-6Y C5,15-methyltransferase (decarboxylating), CbiT subunit (TIGR02469; EC 2.1.1.132; HMM-score: 14.1)Protein synthesis Ribosomal proteins: synthesis and modification protein-(glutamine-N5) methyltransferase, ribosomal protein L3-specific (TIGR03533; EC 2.1.1.-; HMM-score: 13)Protein synthesis tRNA and rRNA base modification ribosomal RNA large subunit methyltransferase J (TIGR00438; EC 2.1.1.166; HMM-score: 11.7)
- TheSEED :
- Sorbitol dehydrogenase homologue (EC:1.1.1.14)
- PFAM: GroES (CL0296) ADH_N; Alcohol dehydrogenase GroES-like domain (PF08240; HMM-score: 96.2)NADP_Rossmann (CL0063) ADH_zinc_N; Zinc-binding dehydrogenase (PF00107; HMM-score: 91)and 9 moreGlu_dehyd_C; Glucose dehydrogenase C-terminus (PF16912; HMM-score: 47.4)AlaDh_PNT_C; Alanine dehydrogenase/PNT, C-terminal domain (PF01262; HMM-score: 23.3)ADH_zinc_N_2; Zinc-binding dehydrogenase (PF13602; HMM-score: 21.4)PrmA; Ribosomal protein L11 methyltransferase (PrmA) (PF06325; HMM-score: 20)Methyltransf_31; Methyltransferase domain (PF13847; HMM-score: 19.4)Methyltransf_25; Methyltransferase domain (PF13649; HMM-score: 17.3)UDPG_MGDP_dh_N; UDP-glucose/GDP-mannose dehydrogenase family, NAD binding domain (PF03721; HMM-score: 15.7)2-Hacid_dh_C; D-isomer specific 2-hydroxyacid dehydrogenase, NAD binding domain (PF02826; HMM-score: 14.3)E-set (CL0159) DUF4179; Domain of unknown function (DUF4179) (PF13786; HMM-score: 13.6)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors: Zn2+
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.67
- Cytoplasmic Membrane Score: 0.01
- Cellwall Score: 0.15
- Extracellular Score: 0.17
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.9314
- Cytoplasmic Membrane Score: 0.028
- Cell wall & surface Score: 0.0004
- Extracellular Score: 0.0402
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.023666
- TAT(Tat/SPI): 0.001679
- LIPO(Sec/SPII): 0.003566
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MKALKLYGVEDLRYEDNEKPVIESANDVIIKVRATGICGSDTSRYKKMGPYIKGMPFGHEFSGVVDAIGSDVTHVNVGDKVTGCPAIPCYQCEYCLKGEYARCEKLFVIGSYEPGSFAEYVKLPAQNVLKVPDNVDYIEAAMVEPSAVVAHGFYKSNIQPGMTVAVMGCGSIGLLAIQWARIFGAAHIIAIDIDAHKLDIATSLGAHQTINSKEENLEKFIENHYANQIDLAIESSGAKVTIGQILTLPKKGGEVVLLGIPYDDIEIDRVHFEKILRNELTVCGSWNCLSSNFPGKEWTATLHYMKTKDINVKPIISHFLPLEKGPETFDKLVNKKERFDKVMFTIY
⊟Experimental data[edit | edit source]
- experimentally validated:
- protein localization:
- quantitative data / protein copy number per cell:
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [1]
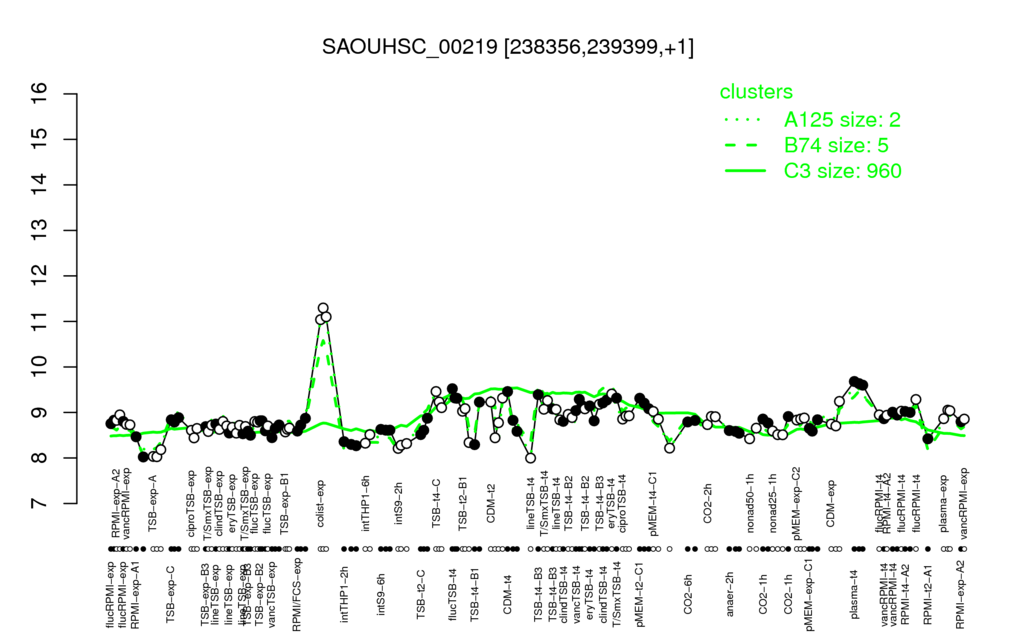 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
