Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_01078
- pan locus tag?: SAUPAN003356000
- symbol: SAOUHSC_01078
- pan gene symbol?: rpmF
- synonym:
- product:
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
⊟Accession numbers[edit | edit source]
- Gene ID: 3919241 NCBI
- RefSeq:
- BioCyc: G1I0R-1014 BioCyc
- MicrobesOnline: 1289538 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121ATGGCAGTACCAAAAAGAAGAACTTCTAAAACTAGAAAAAACAAACGTCGTACGCATTTC
AAAATTTCAGTACCAGGTATGACTGAATGCCCAAACTGTGGCGAATACAAATTATCACAC
CGTGTATGTAAAAACTGTGGTTCTTACAATGGCGAAGAAGTAGCAGCTAAATAA60
120
174
⊟Protein[edit | edit source]
Protein Data Bank: 4WCE
Protein Data Bank: 4WF9
Protein Data Bank: 4WFA
Protein Data Bank: 4WFB
Protein Data Bank: 5HKV
Protein Data Bank: 5HL7
Protein Data Bank: 5LI0
Protein Data Bank: 5ND8
Protein Data Bank: 5ND9
Protein Data Bank: 5NRG
Protein Data Bank: 5TCU
⊟General[edit | edit source]
- locus tag: SAOUHSC_01078
- symbol: SAOUHSC_01078
- description:
- length: 57
- theoretical pI: 10.6639
- theoretical MW: 6484.52
- GRAVY: -1.0614
⊟Function[edit | edit source]
- TIGRFAM: Protein synthesis Ribosomal proteins: synthesis and modification ribosomal protein bL32 (TIGR01031; HMM-score: 97.1)and 3 moreMobile and extrachromosomal element functions Prophage functions phage/conjugal plasmid C-4 type zinc finger protein, TraR family (TIGR02419; HMM-score: 13.3)Regulatory functions DNA interactions putative regulatory protein, FmdB family (TIGR02605; HMM-score: 13.2)YgiT-type zinc finger domain (TIGR03831; HMM-score: 11.9)
- TheSEED :
- LSU ribosomal protein L32p
- LSU ribosomal protein L32p, zinc-dependent
- PFAM: Zn_Beta_Ribbon (CL0167) Ribosomal_L32p; Ribosomal L32p protein family (PF01783; HMM-score: 95.2)and 20 moreZn_ribbon_3; zinc-ribbon domain (PF13248; HMM-score: 24.5)DZR; Double zinc ribbon (PF12773; HMM-score: 18.7)HypA; Hydrogenase/urease nickel incorporation, metallochaperone, hypA (PF01155; HMM-score: 17.6)Zn_Ribbon_1; zinc-ribbon domain (PF13240; HMM-score: 17.3)Nop10p; Nucleolar RNA-binding protein, Nop10p family (PF04135; HMM-score: 15.4)Zn_ribbon_8; Zinc ribbon domain (PF09723; HMM-score: 13.4)no clan defined PolC_DP2_central; DNA polymerase II large subunit DP2, central domain (PF24844; HMM-score: 13.1)Zf_1st_IFT121; IFT121, first zinc finger domain (PF23146; HMM-score: 13)C2H2-zf (CL0361) Znf_XAF1_N; XIAP-associated factor 1-like, N-terminal zinc finger (PF23580; HMM-score: 12.8)Zn_Beta_Ribbon (CL0167) DUF7577; Domain of unknown function (DUF7577) (PF24463; HMM-score: 12.8)Zn_Ribbon_TF; TFIIB zinc-binding (PF08271; HMM-score: 12.6)PFL-like (CL0339) NRDD; Anaerobic ribonucleoside-triphosphate reductase (PF13597; HMM-score: 12.3)no clan defined zf-tcix; Putative treble-clef, zinc-finger, Zn-binding (PF14952; HMM-score: 12.1)Zn_Beta_Ribbon (CL0167) Zn_ribbon_LapB; Rubredoxin metal binding domain (PF18073; HMM-score: 11.9)DUF7575; Domain of unknown function (DUF7575) (PF24460; HMM-score: 11.1)DUF7563; Family of unknown function (DUF7563) (PF24444; HMM-score: 10.7)no clan defined RUBY_RBDX; Rubrerythrin, rubredoxin-like domain (PF21349; HMM-score: 10.4)Zn_Beta_Ribbon (CL0167) Zn_ribbon_IS1595; Transposase zinc-ribbon domain (PF12760; HMM-score: 9.5)zinc_ribbon_6; zinc-ribbon (PF14599; HMM-score: 9.1)Zn_Ribbon_Prim; Zinc-binding domain of primase-helicase (PF08273; HMM-score: 8.2)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 10
- Cytoplasmic Membrane Score: 0
- Cellwall Score: 0
- Extracellular Score: 0
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.6146
- Cytoplasmic Membrane Score: 0
- Cell wall & surface Score: 0
- Extracellular Score: 0.3854
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 0.67
- Signal peptide possibility: -0.5
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.295979
- TAT(Tat/SPI): 0.014651
- LIPO(Sec/SPII): 0.029993
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MAVPKRRTSKTRKNKRRTHFKISVPGMTECPNCGEYKLSHRVCKNCGSYNGEEVAAK
⊟Experimental data[edit | edit source]
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- predicted SigA promoter [3] : SAOUHSC_01077 > S444 > SAOUHSC_01078
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [3]
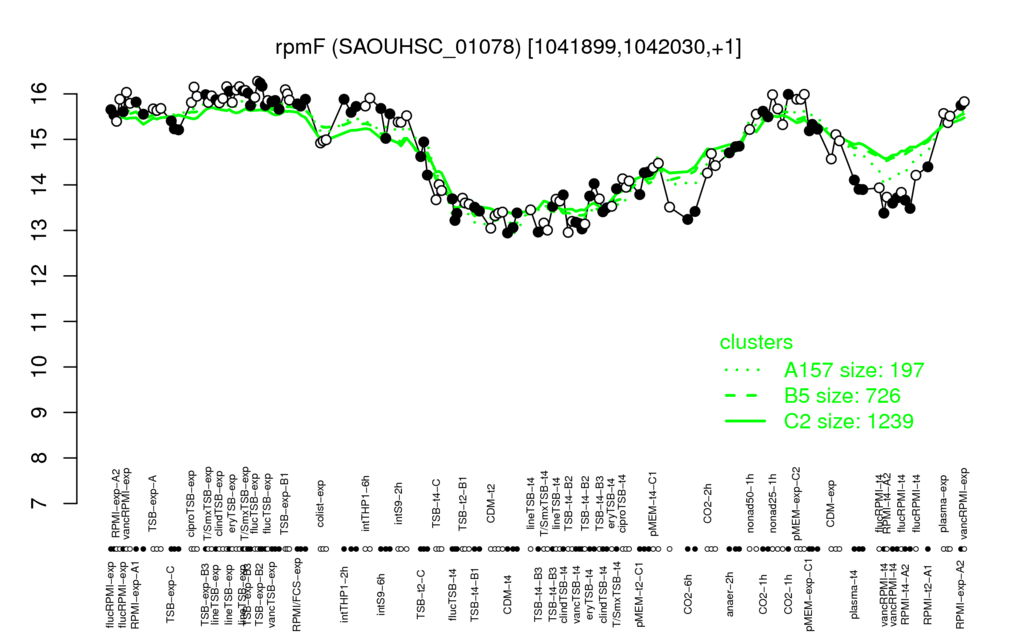 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Roy R Chaudhuri, Andrew G Allen, Paul J Owen, Gil Shalom, Karl Stone, Marcus Harrison, Timothy A Burgis, Michael Lockyer, Jorge Garcia-Lara, Simon J Foster, Stephen J Pleasance, Sarah E Peters, Duncan J Maskell, Ian G Charles
Comprehensive identification of essential Staphylococcus aureus genes using Transposon-Mediated Differential Hybridisation (TMDH).
BMC Genomics: 2009, 10;291
[PubMed:19570206] [WorldCat.org] [DOI] (I e) - ↑ Anne Berscheid, Peter Sass, Konstantin Weber-Lassalle, Ambrose L Cheung, Gabriele Bierbaum
Revisiting the genomes of the Staphylococcus aureus strains NCTC 8325 and RN4220.
Int J Med Microbiol: 2012, 302(2);84-7
[PubMed:22417616] [WorldCat.org] [DOI] (I p) - ↑ 3.0 3.1 3.2 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
