Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_01983
- pan locus tag?: SAUPAN004784000
- symbol: fumC
- pan gene symbol?: fumC
- synonym:
- product: fumarate hydratase
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_01983
- symbol: fumC
- product: fumarate hydratase
- replicon: chromosome
- strand: -
- coordinates: 1887904..1889289
- length: 1386
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3920459 NCBI
- RefSeq: YP_500480 NCBI
- BioCyc: G1I0R-1849 BioCyc
- MicrobesOnline: 1290402 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201
1261
1321
1381ATGTCAGTAAGAATTGAACATGATACTTTTGGAGAAATAGAAGTACCTGCAGATAAATAT
TGGGGTGCTCAAACAGAAAGAAGTAAACGTAATTTCCCAGTTGGTAAAGAGCGTATGCCA
ATCGAAGTAGTTTATGGTTTTGCACAACTAAAGCGTGCAGCAGCAATAGCTAATTTTGAT
TTAGGAAAATTAAGCGAGGCAAAGAAAGATGCCATTGTATACGCATGTGATCAAATTTTA
TCAGGTGAATTAGATGAACACTTCCCACTAGTTGTATGGCAAACAGGAAGCGGTACACAA
AGTAATATGAATGTGAACGAAGTAGTAAGTTATGTTGCTAATATGTATTTAAAAGATCAT
CAAAGTGATGAAAGTATCCACCCAAATGATGATGTAAATAAATCTCAAAGTTCGAATGAT
ACATTCCCAACTGCTATGCACGTTGCATTATATCAAGAGGTTGAAACAAAATTAGAACCT
GCATTAAAACTTTTAAGAAATACTTTGAAAGAAAAAGAAGATAAATTTGATTCAATTATT
AAAATTGGTCGTACACATTTACAAGATGCAACGCCGATCAAACTAGGACAAGAGATTAGT
GGCTGGCGTTATATGCTTGACCGTTGTGAAACAATGTTATCTGAATCTAAGAAGCACATT
TTAAATCTTGCCATCGGTGGTACGGCTGTTGGTACTGGTATTAATGCGCATCCTGAATTT
GGTGATAAAGTGGCACATTATATTTCAGAAAATACGGGTTATCCATTTGTATCTTCTGAA
AATAAATTCCACGCACTTACAGCGCATGATGAAGTTGTTCAATTGCATGGAACATTGAAG
GCATTAGCAGGAGACTTAATGAAAATTGCTAATGATGTGAGATGGTTGGCTTCAGGGCCA
CGAGCTGGTTTGGCAGAAATTTCTATCCCTGAAAATGAACCAGGTTCATCAATTATGCCT
GGTAAAGTTAATCCTACACAATGTGAAATGTTAACAATGGTTGCAGTCCAAGTAATGGGT
AATGATACAGTTGTTGGCTTCGCAAGTTCACAAGGTAACTTTGAATTGAATGTTTATAAA
CCAGTTATTATGCATAATACACTACAATCAATTTATCTTTTAGCTGATGGTATGGAAACA
TTTAATAACAATTGTGCAGTGGGCATTGAACCAATCGAAGAGAATATTGATAATTATTTA
AATCAATCATTAATGTTAGTTACTGCATTAAATCCACATATTGGTTATGAAAAAGCAGCT
CAAATTGCTAAGAAAGCCCATAAAGAAGGTTTAACTTTAAAAGAATCTGCAATTCAAACT
GGATATGTTACAGAAGAACAATTTGAAGCATGGATTAAACCAGAAGATATGGTAGATCCT
CATTAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1260
1320
1380
1386
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_01983
- symbol: FumC
- description: fumarate hydratase
- length: 461
- theoretical pI: 4.97681
- theoretical MW: 51107.5
- GRAVY: -0.329718
⊟Function[edit | edit source]
- reaction: EC 4.2.1.2? ExPASyFumarate hydratase (S)-malate = fumarate + H2O
- TIGRFAM: Energy metabolism TCA cycle fumarate hydratase, class II (TIGR00979; EC 4.2.1.2; HMM-score: 751.7)and 4 moreEnergy metabolism Amino acids and amines aspartate ammonia-lyase (TIGR00839; EC 4.3.1.1; HMM-score: 469.7)Purines, pyrimidines, nucleosides, and nucleotides Purine ribonucleotide biosynthesis adenylosuccinate lyase (TIGR00928; EC 4.3.2.2; HMM-score: 88.9)Energy metabolism Other 3-carboxy-cis,cis-muconate cycloisomerase (TIGR02426; EC 5.5.1.2; HMM-score: 73)Amino acid biosynthesis Glutamate family argininosuccinate lyase (TIGR00838; EC 4.3.2.1; HMM-score: 60.3)
- TheSEED :
- Fumarate hydratase class II (EC 4.2.1.2)
- PFAM: no clan defined Lyase_1; Lyase (PF00206; HMM-score: 379.6)and 5 moreFumaraseC_C; Fumarase C C-terminus (PF10415; HMM-score: 90.6)DNA_primase_lrg (CL0242) DNA_primase_lrg; Eukaryotic and archaeal DNA primase, large subunit (PF04104; HMM-score: 16.9)no clan defined Rubis-subs-bind; Rubisco LSMT substrate-binding (PF09273; HMM-score: 14.9)P-loop_NTPase (CL0023) G-alpha; G-protein alpha subunit (PF00503; HMM-score: 13.9)KNTase_C (CL0291) NTase_sub_bind; Nucleotidyltransferase substrate binding protein like (PF08780; HMM-score: 12.3)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.97
- Cytoplasmic Membrane Score: 0
- Cellwall Score: 0.01
- Extracellular Score: 0.02
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.9369
- Cytoplasmic Membrane Score: 0.0008
- Cell wall & surface Score: 0.0001
- Extracellular Score: 0.0622
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.003889
- TAT(Tat/SPI): 0.000959
- LIPO(Sec/SPII): 0.000775
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MSVRIEHDTFGEIEVPADKYWGAQTERSKRNFPVGKERMPIEVVYGFAQLKRAAAIANFDLGKLSEAKKDAIVYACDQILSGELDEHFPLVVWQTGSGTQSNMNVNEVVSYVANMYLKDHQSDESIHPNDDVNKSQSSNDTFPTAMHVALYQEVETKLEPALKLLRNTLKEKEDKFDSIIKIGRTHLQDATPIKLGQEISGWRYMLDRCETMLSESKKHILNLAIGGTAVGTGINAHPEFGDKVAHYISENTGYPFVSSENKFHALTAHDEVVQLHGTLKALAGDLMKIANDVRWLASGPRAGLAEISIPENEPGSSIMPGKVNPTQCEMLTMVAVQVMGNDTVVGFASSQGNFELNVYKPVIMHNTLQSIYLLADGMETFNNNCAVGIEPIEENIDNYLNQSLMLVTALNPHIGYEKAAQIAKKAHKEGLTLKESAIQTGYVTEEQFEAWIKPEDMVDPH
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas [1] [2]
- protein localization: data available for COL
- quantitative data / protein copy number per cell: data available for COL
- interaction partners:
SAOUHSC_02490 (adk) adenylate kinase [3] (data from MRSA252) SAOUHSC_01683 (dnaK) molecular chaperone DnaK [3] (data from MRSA252) SAOUHSC_00574 (eutD) phosphotransacetylase [3] (data from MRSA252) SAOUHSC_02117 (gatA) aspartyl/glutamyl-tRNA amidotransferase subunit A [3] (data from MRSA252) SAOUHSC_00900 (pgi) glucose-6-phosphate isomerase [3] (data from MRSA252) SAOUHSC_00520 (rplJ) 50S ribosomal protein L10 [3] (data from MRSA252) SAOUHSC_02492 (rplO) 50S ribosomal protein L15 [3] (data from MRSA252) SAOUHSC_02484 (rplQ) 50S ribosomal protein L17 [3] (data from MRSA252) SAOUHSC_02510 (rplW) 50S ribosomal protein L23 [3] (data from MRSA252) SAOUHSC_00524 (rpoB) DNA-directed RNA polymerase subunit beta [3] (data from MRSA252) SAOUHSC_01232 (rpsB) 30S ribosomal protein S2 [3] (data from MRSA252) SAOUHSC_02494 (rpsE) 30S ribosomal protein S5 [3] (data from MRSA252) SAOUHSC_02353 (upp) uracil phosphoribosyltransferase [3] (data from MRSA252) SAOUHSC_00336 acetyl-CoA acyltransferase [3] (data from MRSA252) SAOUHSC_00529 elongation factor G [3] (data from MRSA252) SAOUHSC_00669 hypothetical protein [3] (data from MRSA252) SAOUHSC_00675 hypothetical protein [3] (data from MRSA252) SAOUHSC_00679 hypothetical protein [3] (data from MRSA252) SAOUHSC_00795 glyceraldehyde-3-phosphate dehydrogenase [3] (data from MRSA252) SAOUHSC_00878 hypothetical protein [3] (data from MRSA252) SAOUHSC_00895 glutamate dehydrogenase [3] (data from MRSA252) SAOUHSC_00951 hypothetical protein [3] (data from MRSA252) SAOUHSC_01040 pyruvate dehydrogenase complex, E1 component subunit alpha [3] (data from MRSA252) SAOUHSC_01801 isocitrate dehydrogenase [3] (data from MRSA252)
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: no polycistronic organisation predicted
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [4]
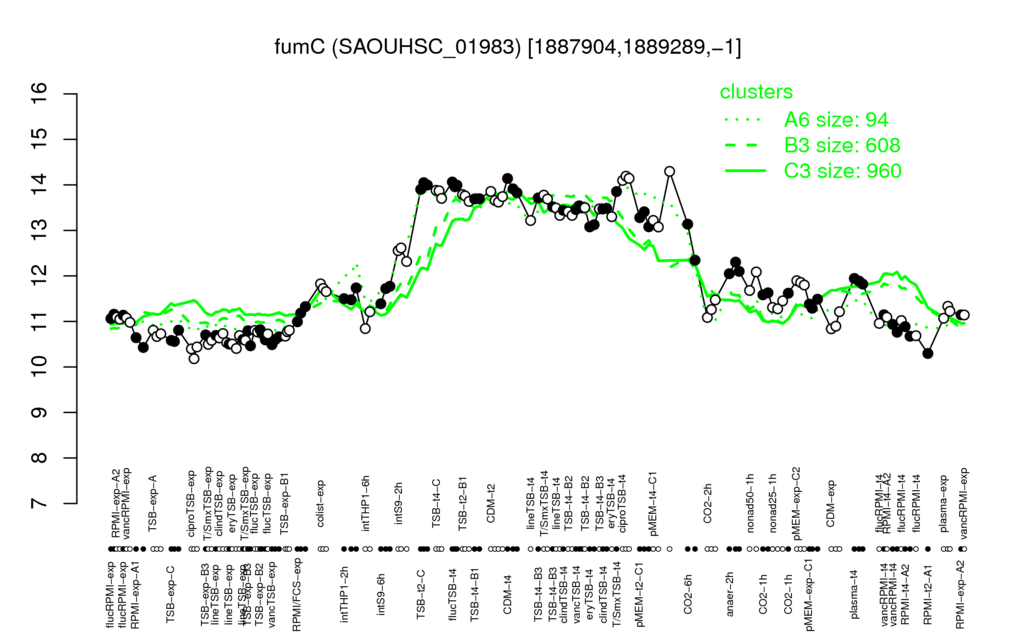 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Maren Depke, Stephan Michalik, Alexander Rabe, Kristin Surmann, Lars Brinkmann, Nico Jehmlich, Jörg Bernhardt, Michael Hecker, Bernd Wollscheid, Zhi Sun, Robert L Moritz, Uwe Völker, Frank Schmidt
A peptide resource for the analysis of Staphylococcus aureus in host-pathogen interaction studies.
Proteomics: 2015, 15(21);3648-61
[PubMed:26224020] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Maren Depke, Annette Murr, Manuela Gesell Salazar, Ulrike Kusebauch, Zhi Sun, Tanja C Meyer, Kristin Surmann, Henrike Pförtner, Petra Hildebrandt, Stefan Weiss, Laura Marcela Palma Medina, Melanie Gutjahr, Elke Hammer, Dörte Becher, Thomas Pribyl, Sven Hammerschmidt, Eric W Deutsch, Samuel L Bader, Michael Hecker, Robert L Moritz, Ulrike Mäder, Uwe Völker, Frank Schmidt
A global Staphylococcus aureus proteome resource applied to the in vivo characterization of host-pathogen interactions.
Sci Rep: 2017, 7(1);9718
[PubMed:28887440] [WorldCat.org] [DOI] (I e) - ↑ 3.00 3.01 3.02 3.03 3.04 3.05 3.06 3.07 3.08 3.09 3.10 3.11 3.12 3.13 3.14 3.15 3.16 3.17 3.18 3.19 3.20 3.21 3.22 3.23 Artem Cherkasov, Michael Hsing, Roya Zoraghi, Leonard J Foster, Raymond H See, Nikolay Stoynov, Jihong Jiang, Sukhbir Kaur, Tian Lian, Linda Jackson, Huansheng Gong, Rick Swayze, Emily Amandoron, Farhad Hormozdiari, Phuong Dao, Cenk Sahinalp, Osvaldo Santos-Filho, Peter Axerio-Cilies, Kendall Byler, William R McMaster, Robert C Brunham, B Brett Finlay, Neil E Reiner
Mapping the protein interaction network in methicillin-resistant Staphylococcus aureus.
J Proteome Res: 2011, 10(3);1139-50
[PubMed:21166474] [WorldCat.org] [DOI] (I p) - ↑ Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
