Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_00668
- pan locus tag?: SAUPAN002541000
- symbol: SAOUHSC_00668
- pan gene symbol?: vraG
- synonym:
- product: ABC transporter permease
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_00668
- symbol: SAOUHSC_00668
- product: ABC transporter permease
- replicon: chromosome
- strand: +
- coordinates: 654690..656579
- length: 1890
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3919431 NCBI
- RefSeq: YP_499227 NCBI
- BioCyc: G1I0R-622 BioCyc
- MicrobesOnline: 1289137 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201
1261
1321
1381
1441
1501
1561
1621
1681
1741
1801
1861ATGACCTTTAACGAGATAATATTTAAAAATTTCCGTCAAAATTTATCACATTATGCCATC
TATCTTTTTTCGTTAATTACGAGTGTAGTATTGTATTTTAGCTTTGTAGCATTAAAATAC
GCTCATAAACTAAACATGACAGAGTCATATCCAATTATAAAGGAAGGCTCACAAGTCGGA
AGCTACTTTCTATTTTTCATCATAATTGCATTTTTGTTATATGCCAATGTGTTATTTATT
AAACGACGAAGTTATGAGCTTGCATTATATCAAACATTAGGTTTATCTAAATTCAACATT
ATTTATATACTAATGCTCGAACAATTACTAATATTTATAATTACGGCAATATTAGGTATT
ATTATTGGTATTTTTGGTTCGAAACTGTTATTAATGATTGTCTTTACATTATTAGGAATT
AAAGAAAAGGTTCCAATTATTTTTAGTTTGAGGGCGGTATTTGAAACATTAATGTTAATC
GGTGTCGCTTATTTTTTAACATCTGCTCAAAATTTTATATTAGTGTTCAAACAATCTATT
TCACAGATGTCAAAGAATAACCAGGTTAAAGAAACAAATCATAATAAAATTACATTTGAA
GAGGTTGTTTTAGGCATCTTAGGTATAGTATTGATTACCACAGGATACTATCTATCTTTG
AACATTGTTCAATATTATGATTCTATCGGTACACTTATGTTTATTTTATTGTCAACTGTG
ATTGGGGCATACTTATTTTTTAAAAGCTCTGTTTCTCTAGTTTTTAAAATGGTGAAGAAG
TTTAGAAAAGGTGTTATAAGTGTAAATGATGTCATGTTCTCATCATCTATTATGTATCGT
ATTAAGAAAAATGCTTTTTCACTTACGGTCATGGCAATCATTTCAGCGATTACTGTTTCA
GTTCTTTGCTTTGCTGCTATAAGTAGAGCGTCCTTATCAAGTGAAATAAAATATACTGCA
CCACACGACGTTACAATTAAAGACCAACAAAAAGCTAATCAATTAGCAAGTGAATTAAAC
AATCAAAAAATTCCTCATTTTTATAATTATAAAGAAGTAATTCATACGAAATTGTATAAA
GATAATTTATTTGATGTAAAAGCGAAAGAACCATACAATGTAACAATTACTAGTGATAAA
TACATCCCTAATACTGATTTGAAACGTGGGCAAGCTGATTTATTTGTAGCGGAAGGTTCT
ATCAAAGATTTAGTGAAACATAAGAAGCATGGTAAGGCAATTATAGGAACGAAAAAACAT
CATGTTAATATTAAGTTACGTAAAGATATTAATAAAATCTATTTTATGACAGATGTTGAT
TTAGGTGGACCAACGTTTGTCTTAAATGACAAAGACTATCAAGAAATAAGAAAGTATACA
AAGGCAAAGCATATCGTCTCTCAATTTGGATTCGATTTGAAACATAAAAAAGATGCTTTA
GCATTAGAAAAAGCGAAAAATAAAGTTGATAAATCTATTGAAACAAGAAGTGAAGCGATA
AGCTCAATATCAAGTTTAACCGGAATATTATTATTTGTAACATCATTTTTAGGTATTACA
TTCTTGATTGCTGTATGTTGCATTATATACATAAAGCAAATAGATGAAACCGAAGATGAG
TTAGAGAATTATAGTATTTTGAGAAAGCTTGGATTTACACAAAAAGATATGGCAAGGGGA
CTAAAGTTTAAAATTATGTTTAATTTTGGGTTACCTTTAGTTATTGCACTATCACATGCA
TATTTTACATCATTAGCATATATGAAATTAATGGGTACAACGAATCAAATACCGGTTTTC
ATAGTAATGGGATTATACATTTGTATGTATGCTGTTTTTGCAGTGACGGCTTATAATCAT
TCCAAGCGAACAATTAGACATTCCATATAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1260
1320
1380
1440
1500
1560
1620
1680
1740
1800
1860
1890
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_00668
- symbol: SAOUHSC_00668
- description: ABC transporter permease
- length: 629
- theoretical pI: 10.0671
- theoretical MW: 71731.5
- GRAVY: 0.355008
⊟Function[edit | edit source]
- TIGRFAM: acidobacterial duplicated orphan permease (TIGR03434; HMM-score: 13.9)
- TheSEED :
- Bacitracin export permease protein BceB
- PFAM: BPD_transp_1 (CL0404) FtsX; FtsX-like permease C-terminal (PF02687; HMM-score: 34.1)and 2 moreno clan defined DUF6518; Family of unknown function (DUF6518) (PF20128; HMM-score: 13.7)FxsA; FxsA cytoplasmic membrane protein (PF04186; HMM-score: 8.2)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic Membrane
- Cytoplasmic Score: 0
- Cytoplasmic Membrane Score: 10
- Cellwall Score: 0
- Extracellular Score: 0
- Internal Helices: 10
- DeepLocPro: Cytoplasmic Membrane
- Cytoplasmic Score: 0
- Cytoplasmic Membrane Score: 0.9999
- Cell wall & surface Score: 0
- Extracellular Score: 0.0001
- LocateP: Multi-transmembrane
- Prediction by SwissProt Classification: Membrane
- Pathway Prediction: Sec-(SPI)
- Intracellular possibility: 0.17
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.007787
- TAT(Tat/SPI): 0.000247
- LIPO(Sec/SPII): 0.002804
- predicted transmembrane helices (TMHMM): 10
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MTFNEIIFKNFRQNLSHYAIYLFSLITSVVLYFSFVALKYAHKLNMTESYPIIKEGSQVGSYFLFFIIIAFLLYANVLFIKRRSYELALYQTLGLSKFNIIYILMLEQLLIFIITAILGIIIGIFGSKLLLMIVFTLLGIKEKVPIIFSLRAVFETLMLIGVAYFLTSAQNFILVFKQSISQMSKNNQVKETNHNKITFEEVVLGILGIVLITTGYYLSLNIVQYYDSIGTLMFILLSTVIGAYLFFKSSVSLVFKMVKKFRKGVISVNDVMFSSSIMYRIKKNAFSLTVMAIISAITVSVLCFAAISRASLSSEIKYTAPHDVTIKDQQKANQLASELNNQKIPHFYNYKEVIHTKLYKDNLFDVKAKEPYNVTITSDKYIPNTDLKRGQADLFVAEGSIKDLVKHKKHGKAIIGTKKHHVNIKLRKDINKIYFMTDVDLGGPTFVLNDKDYQEIRKYTKAKHIVSQFGFDLKHKKDALALEKAKNKVDKSIETRSEAISSISSLTGILLFVTSFLGITFLIAVCCIIYIKQIDETEDELENYSILRKLGFTQKDMARGLKFKIMFNFGLPLVIALSHAYFTSLAYMKLMGTTNQIPVFIVMGLYICMYAVFAVTAYNHSKRTIRHSI
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas [1] [2]
- protein localization: data available for COL
- quantitative data / protein copy number per cell:
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: SAOUHSC_00664 > SAOUHSC_00665 > SAOUHSC_00666 > SAOUHSC_00667 > SAOUHSC_00668predicted SigA promoter [3] : S251 > SAOUHSC_00664 > SAOUHSC_00665 > SAOUHSC_00666 > S252 > SAOUHSC_00667 > SAOUHSC_00668
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [3]
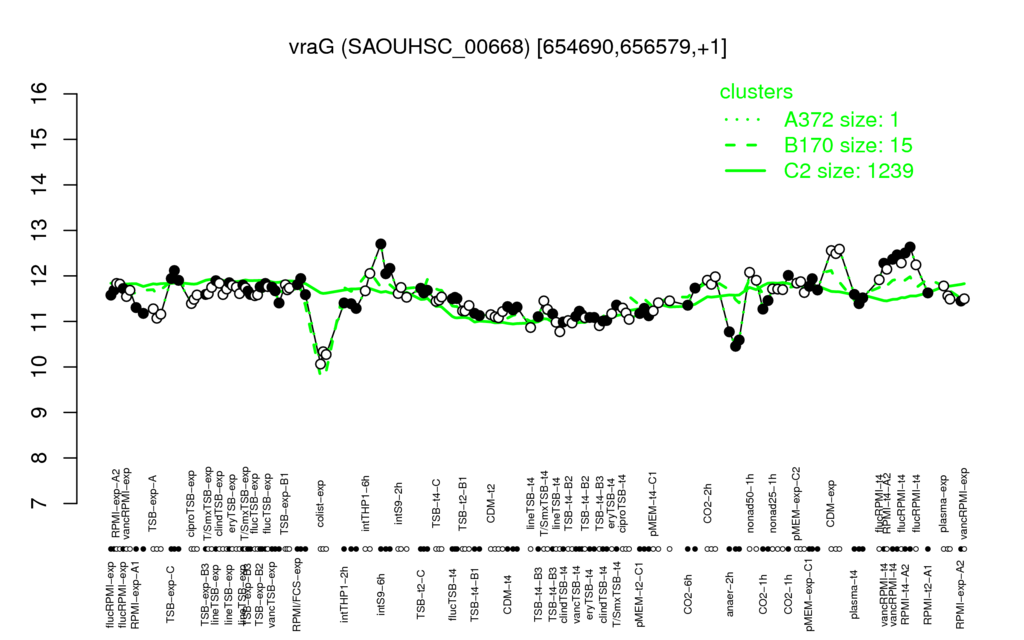 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Maren Depke, Stephan Michalik, Alexander Rabe, Kristin Surmann, Lars Brinkmann, Nico Jehmlich, Jörg Bernhardt, Michael Hecker, Bernd Wollscheid, Zhi Sun, Robert L Moritz, Uwe Völker, Frank Schmidt
A peptide resource for the analysis of Staphylococcus aureus in host-pathogen interaction studies.
Proteomics: 2015, 15(21);3648-61
[PubMed:26224020] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Maren Depke, Annette Murr, Manuela Gesell Salazar, Ulrike Kusebauch, Zhi Sun, Tanja C Meyer, Kristin Surmann, Henrike Pförtner, Petra Hildebrandt, Stefan Weiss, Laura Marcela Palma Medina, Melanie Gutjahr, Elke Hammer, Dörte Becher, Thomas Pribyl, Sven Hammerschmidt, Eric W Deutsch, Samuel L Bader, Michael Hecker, Robert L Moritz, Ulrike Mäder, Uwe Völker, Frank Schmidt
A global Staphylococcus aureus proteome resource applied to the in vivo characterization of host-pathogen interactions.
Sci Rep: 2017, 7(1);9718
[PubMed:28887440] [WorldCat.org] [DOI] (I e) - ↑ 3.0 3.1 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
