Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_01949
- pan locus tag?: SAUPAN004575000
- symbol: SAOUHSC_01949
- pan gene symbol?: epiP
- synonym:
- product: intracellular serine protease
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_01949
- symbol: SAOUHSC_01949
- product: intracellular serine protease
- replicon: chromosome
- strand: -
- coordinates: 1851533..1852906
- length: 1374
- essential: no DEG
⊟Accession numbers[edit | edit source]
- Gene ID: 3920894 NCBI
- RefSeq: YP_500448 NCBI
- BioCyc: G1I0R-1810 BioCyc
- MicrobesOnline: 1290362 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201
1261
1321ATGAAAATCATAAAGCGCGCTATTATTAGTTTAATAATATTGAGTCTTTTAATATCTATT
ACAATGAGTAATGCATCAGCGTCAGAAGAACTATATTACAGTGTTGAATACAAAAATACA
GCAACATTTAACAAGTTAGTTAAAAAGAAGTCCTTAAACGTTGTCTATAATATACCGGAA
TTACATGTGGCACAGATTAAAATGACGAAAATGCATGCTAATGCTTTAGCAAACTATAAA
AATGATATTAAATATATCAATGCCACATGTTCAACTTGTATTACTAGCGAGAAAACAATA
GACAGAACATCAAATGAGTCATTATTTTCAAGACAATGGGATATGAATAAAATAACCAAT
AATGGTGCATCGTATGATGATTTGCCAAAACATGCTAACACCAAAATAGCAATCATAGAT
ACAGGTGTGATGAAAAACCATGACGATTTGAAAAATAATTTCTCGACTGATTCTAAAAAT
TTAGTACCTTTAAACGGTTTTAGAGGTACTGAACCGGAGGAAACAGGTGATGTTCACGAT
GTCAATGATAGGAAAGGACATGGCACGATGGTGTCGGGTCAAACGAGTGCTAATGGTAAG
TTAATAGGTGTTGCACCGAATAACAAATTTACAATGTATCGCGTGTTTGGTAGTAAAAAA
ACAGAACTGCTTTGGGTATCAAAAGCGATTGTTCAAGCTGCAAATGATGGAAATCAAGTC
ATTAATATTAGTGTTGGTAGTTATATTATTTTGGACAAAAATGACCATCAAACATTTAGA
AAAGATGAAAAAGTAGAATACGATGCGTTACAGAAAGCAATCAATTACGCCAAGAAGAAA
AAATCTATCGTTGTTGCTGCAGCTGGTAATGATGGTATTGATGTCAATGACAAACAGAAA
CTAAAATTACAGCGTGAATATCAAGGTAATGGCGAAGTGAAAGATGTTCCTGCATCTATG
GACAATGTCGTTACAGTAGGATCTACAGATCAAAAGAGTAATCTATCTGAGTTTTCCAAT
TTTGGTATGAATTATACAGATATTGCTGCGCCCGGAGGATCATTTGCTTATTTAAATCAA
TTCGGTGTGGATAAATGGATGAATGAAGGGTATATGCATAAGGAGAACATTTTAACTACT
GCCAATAACGGAAGATATATTTATCAAGCTGGAACTTCATTAGCCACACCTAAAGTTTCG
GGAGCACTAGCTTTAATCATTGATAAATATCATCTTGAAAAACATCCAGATAAAGCGATT
GAATTGTTATATCAGCATGGGACATCTAAGAATAATAAACCATTTAGTAGATATGGGCAT
GGTGAGCTTGATGTGTATAAAGCATTAAATGTAGCAAATCAAAAAGCAAGTTAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1260
1320
1374
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_01949
- symbol: SAOUHSC_01949
- description: intracellular serine protease
- length: 457
- theoretical pI: 9.48761
- theoretical MW: 50735.1
- GRAVY: -0.497812
⊟Function[edit | edit source]
- TIGRFAM: Protein fate Protein and peptide secretion and trafficking type VII secretion-associated serine protease mycosin (TIGR03921; EC 3.4.21.-; HMM-score: 136)Protein fate Protein modification and repair type VII secretion-associated serine protease mycosin (TIGR03921; EC 3.4.21.-; HMM-score: 136)and 1 morecyanobactin maturation protease, PatA/PatG family (TIGR03895; EC 3.4.21.-; HMM-score: 50.3)
- TheSEED :
- Epidermin leader peptide processing serine protease epiP precursor (EC 3.4.21.-)
- PFAM: no clan defined Peptidase_S8; Subtilase family (PF00082; HMM-score: 176.9)PPP-I (CL0570) PPI; Protease propeptides/inhibitors (PF22386; HMM-score: 151.4)and 2 moreno clan defined Cpf1_PI-like; CRISPR-associated endonuclease Cpf1 PI domain (PF22222; HMM-score: 14.5)DUF1514; Protein of unknown function (DUF1514) (PF07438; HMM-score: 12)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Extracellular
- Cytoplasmic Score: 0
- Cytoplasmic Membrane Score: 0.09
- Cellwall Score: 0.18
- Extracellular Score: 9.73
- Internal Helices: 0
- DeepLocPro: Extracellular
- Cytoplasmic Score: 0
- Cytoplasmic Membrane Score: 0.0037
- Cell wall & surface Score: 0.0045
- Extracellular Score: 0.9917
- LocateP: Secretory(released) (with CS)
- Prediction by SwissProt Classification: Extracellular
- Pathway Prediction: Sec-(SPI)
- Intracellular possibility: 0.17
- Signal peptide possibility: 1
- N-terminally Anchored Score: -1
- Predicted Cleavage Site: SNASASEE
- SignalP: Signal peptide SP(Sec/SPI) length 27 aa
- SP(Sec/SPI): 0.997615
- TAT(Tat/SPI): 0.000491
- LIPO(Sec/SPII): 0.001085
- Cleavage Site: CS pos: 27-28. ASA-SE. Pr: 0.9602
- predicted transmembrane helices (TMHMM): 1
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MKIIKRAIISLIILSLLISITMSNASASEELYYSVEYKNTATFNKLVKKKSLNVVYNIPELHVAQIKMTKMHANALANYKNDIKYINATCSTCITSEKTIDRTSNESLFSRQWDMNKITNNGASYDDLPKHANTKIAIIDTGVMKNHDDLKNNFSTDSKNLVPLNGFRGTEPEETGDVHDVNDRKGHGTMVSGQTSANGKLIGVAPNNKFTMYRVFGSKKTELLWVSKAIVQAANDGNQVINISVGSYIILDKNDHQTFRKDEKVEYDALQKAINYAKKKKSIVVAAAGNDGIDVNDKQKLKLQREYQGNGEVKDVPASMDNVVTVGSTDQKSNLSEFSNFGMNYTDIAAPGGSFAYLNQFGVDKWMNEGYMHKENILTTANNGRYIYQAGTSLATPKVSGALALIIDKYHLEKHPDKAIELLYQHGTSKNNKPFSRYGHGELDVYKALNVANQKAS
⊟Experimental data[edit | edit source]
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [1]
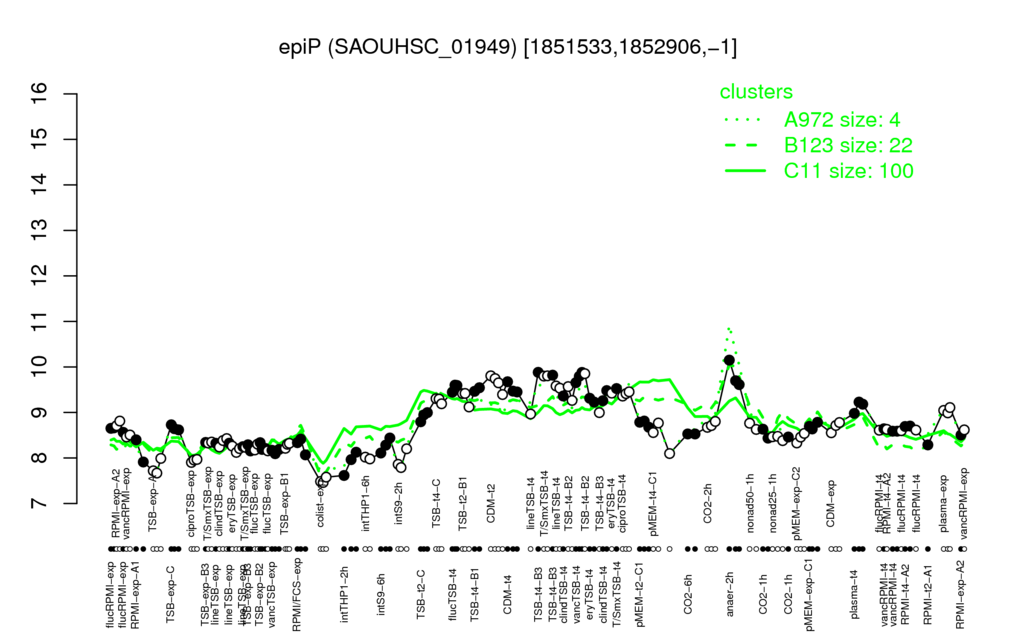 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
