Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_00341
- pan locus tag?: SAUPAN001903000
- symbol: SAOUHSC_00341
- pan gene symbol?: metI
- synonym:
- product: hypothetical protein
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_00341
- symbol: SAOUHSC_00341
- product: hypothetical protein
- replicon: chromosome
- strand: -
- coordinates: 355117..356220
- length: 1104
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3921815 NCBI
- RefSeq: YP_498931 NCBI
- BioCyc: G1I0R-317 BioCyc
- MicrobesOnline: 1288825 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081ATGAAGGATACACAGTTAGCCCAAATCACATTAACCGATGATTCAACCGGTGCTATAGCG
AATCCAATCCATTTATCTACTGCCTACAAGCATCCAAAACTAGGACAATCGACAGGTTTT
GATTATACACGTACTAAAAATCCTACACGCTCAACATTTGAAACCTGTTTTGCCAAACTT
GAGCATGGTATTGCATCATTCGCTACATCAAGTGGAATGTCAGCCATTCAATTAATATGT
AATCTATTTAAACCTCATGATGAAATTTTAGTTTCATTCGATTTATATGGTGGCACATTT
AGATTATTTGAATTTTACGAGCAACAATACGATATCAAATTTAAGTACGTTGATTTTACA
GATTATGAACAAGTTGAAAAAGAAATCACTGATAAAACAGTTGCATTATTCATTGAACCA
ATATCTAACCCACAAATGATTGCTATTGATGTAAAGCCATACTATCAACTTTGTAAAGCT
AAAGGCTTATTGTCAATTATCGACAATACTTTTTTAACACCTTATCTTTCAACACCACTA
GCAGAAGGTGCTGATATAGTCTTACATTCAGCCACGAAATATATTGGCGGACATAACGAT
GTACTAGCAGGTGTCGTAACCGTCAAAGATGAATCACTCGCGCAACAGTTGTTTGATTTT
CACAACATGACTGGCGCAACACTTTCACCAATAGATAGTTATTTGTTGTTACGTGGACTT
AAAACTTTGCATTTACGCATTGAGCGTGCGCAATCAAACGCTAGAAAACTTGCTAAAAAA
TGTCAGTCACTTCAAGCAATTGACGAAGTACTATATAGCGGGCAAACTGGCATGCTTAGT
TTAAGACTTAACAAGGCCTATAGCGTCGCTAAATTATTAGAAAATTTAGACATTTGCATT
TTTGCAGAAAGTTTAGGAGGTACTGAAACATTAGTGACCTTCCCTTACACCCAAACACAT
GTTGATATGCCAGATGCTGAAAAAGATAAACGTGGCATTGATGAGTATTTAATCCGCTTA
TCACTAGGTGTTGAAAATTATGAAGACATCGAACGCGATATCATCCAAGCATTAGATAAA
GCTCAGATTGGAGAGATTGTATGA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1104
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_00341
- symbol: SAOUHSC_00341
- description: hypothetical protein
- length: 367
- theoretical pI: 5.01375
- theoretical MW: 41071.6
- GRAVY: -0.134332
⊟Function[edit | edit source]
- TIGRFAM: Amino acid biosynthesis Aspartate family O-succinylhomoserine (thiol)-lyase (TIGR02080; EC 2.5.1.48; HMM-score: 312.8)Energy metabolism Amino acids and amines methionine gamma-lyase (TIGR01328; EC 4.4.1.11; HMM-score: 280.4)cystathionine beta-lyase (TIGR01329; EC 4.4.1.8; HMM-score: 277.7)and 8 moreAmino acid biosynthesis Aspartate family O-acetylhomoserine aminocarboxypropyltransferase/cysteine synthase (TIGR01326; HMM-score: 241.8)Amino acid biosynthesis Serine family O-acetylhomoserine aminocarboxypropyltransferase/cysteine synthase (TIGR01326; HMM-score: 241.8)Amino acid biosynthesis Aspartate family O-succinylhomoserine sulfhydrylase (TIGR01325; EC 4.2.99.-; HMM-score: 237.2)Amino acid biosynthesis Aspartate family cystathionine beta-lyase (TIGR01324; EC 4.4.1.8; HMM-score: 129.3)transaminase, acetylornithine/succinylornithine family (TIGR00707; HMM-score: 17.7)Unknown function Enzymes of unknown specificity cysteine desulfurase family protein (TIGR01977; HMM-score: 17.7)Unknown function Enzymes of unknown specificity uncharacterized pyridoxal phosphate-dependent enzyme (TIGR01437; HMM-score: 14.3)Energy metabolism Amino acids and amines succinylornithine transaminase family (TIGR03246; EC 2.6.1.81; HMM-score: 12.6)
- TheSEED :
- Cystathionine gamma-synthase (EC 2.5.1.48)
Amino Acids and Derivatives Lysine, threonine, methionine, and cysteine Methionine Biosynthesis Cystathionine gamma-synthase (EC 2.5.1.48)and 1 more - PFAM: PLP_aminotran (CL0061) Cys_Met_Meta_PP; Cys/Met metabolism PLP-dependent enzyme (PF01053; HMM-score: 376.2)and 3 moreAminotran_1_2; Aminotransferase class I and II (PF00155; HMM-score: 21.2)Aminotran_5; Aminotransferase class-V (PF00266; HMM-score: 19.5)Beta_propeller (CL0186) Rod_N; Rod, N-terminal (PF24504; HMM-score: 12.5)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors: pyridoxal 5'-phosphate
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.97
- Cytoplasmic Membrane Score: 0
- Cellwall Score: 0.01
- Extracellular Score: 0.02
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.9876
- Cytoplasmic Membrane Score: 0.001
- Cell wall & surface Score: 0.0001
- Extracellular Score: 0.0112
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.032735
- TAT(Tat/SPI): 0.002987
- LIPO(Sec/SPII): 0.002616
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MKDTQLAQITLTDDSTGAIANPIHLSTAYKHPKLGQSTGFDYTRTKNPTRSTFETCFAKLEHGIASFATSSGMSAIQLICNLFKPHDEILVSFDLYGGTFRLFEFYEQQYDIKFKYVDFTDYEQVEKEITDKTVALFIEPISNPQMIAIDVKPYYQLCKAKGLLSIIDNTFLTPYLSTPLAEGADIVLHSATKYIGGHNDVLAGVVTVKDESLAQQLFDFHNMTGATLSPIDSYLLLRGLKTLHLRIERAQSNARKLAKKCQSLQAIDEVLYSGQTGMLSLRLNKAYSVAKLLENLDICIFAESLGGTETLVTFPYTQTHVDMPDAEKDKRGIDEYLIRLSLGVENYEDIERDIIQALDKAQIGEIV
⊟Experimental data[edit | edit source]
- experimentally validated: data available for COL
- protein localization: data available for COL
- quantitative data / protein copy number per cell: data available for COL
- interaction partners:
SAOUHSC_01683 (dnaK) molecular chaperone DnaK [1] (data from MRSA252) SAOUHSC_00799 (eno) phosphopyruvate hydratase [1] (data from MRSA252) SAOUHSC_02496 (rplF) 50S ribosomal protein L6 [1] (data from MRSA252) SAOUHSC_00521 (rplL) 50S ribosomal protein L7/L12 [1] (data from MRSA252) SAOUHSC_01784 (rplT) 50S ribosomal protein L20 [1] (data from MRSA252) SAOUHSC_01757 (rplU) 50S ribosomal protein L21 [1] (data from MRSA252) SAOUHSC_02477 (rpsI) 30S ribosomal protein S9 [1] (data from MRSA252) SAOUHSC_01234 (tsf) elongation factor Ts [1] (data from MRSA252) SAOUHSC_00187 formate acetyltransferase [1] (data from MRSA252) SAOUHSC_00530 elongation factor Tu [1] (data from MRSA252) SAOUHSC_01040 pyruvate dehydrogenase complex, E1 component subunit alpha [1] (data from MRSA252) SAOUHSC_01041 pyruvate dehydrogenase complex, E1 component subunit beta [1] (data from MRSA252) SAOUHSC_01042 branched-chain alpha-keto acid dehydrogenase subunit E2 [1] (data from MRSA252) SAOUHSC_01806 pyruvate kinase [1] (data from MRSA252)
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: SAOUHSC_00338 < SAOUHSC_00339 < SAOUHSC_00340 < SAOUHSC_00341predicted SigB promoter [2] : SAOUHSC_00337 < SAOUHSC_00338 < SAOUHSC_00339 < SAOUHSC_00340 < SAOUHSC_00341
⊟Regulation[edit | edit source]
- regulator: T-box(Met) (transcription antitermination) regulon
T-box(Met) (5' cis-acting region) important in Amino acid metabolism; compare RegPrecise for N315
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [2]
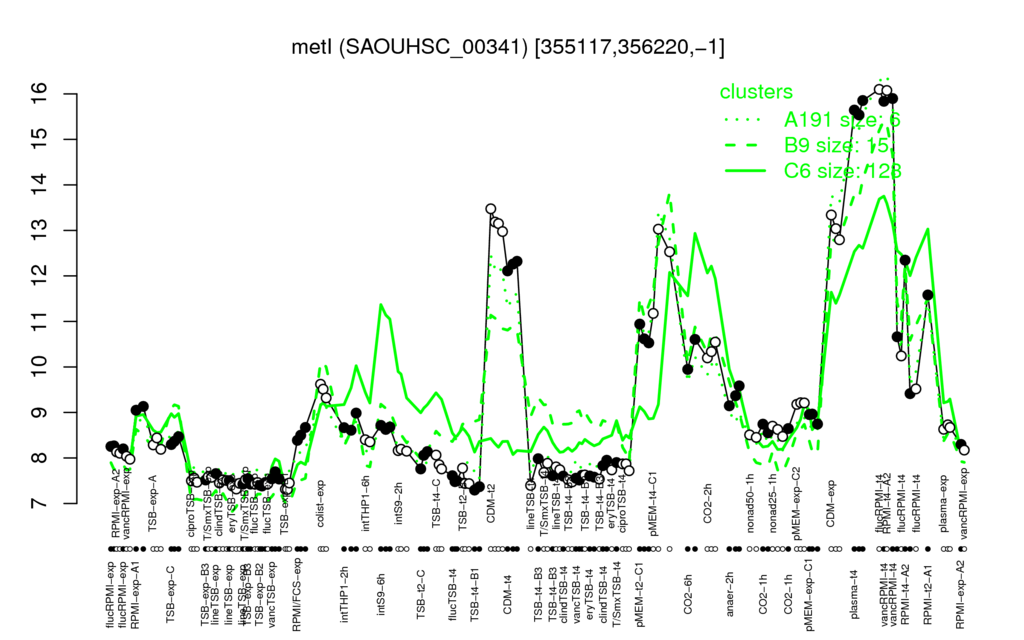 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 1.13 Artem Cherkasov, Michael Hsing, Roya Zoraghi, Leonard J Foster, Raymond H See, Nikolay Stoynov, Jihong Jiang, Sukhbir Kaur, Tian Lian, Linda Jackson, Huansheng Gong, Rick Swayze, Emily Amandoron, Farhad Hormozdiari, Phuong Dao, Cenk Sahinalp, Osvaldo Santos-Filho, Peter Axerio-Cilies, Kendall Byler, William R McMaster, Robert C Brunham, B Brett Finlay, Neil E Reiner
Mapping the protein interaction network in methicillin-resistant Staphylococcus aureus.
J Proteome Res: 2011, 10(3);1139-50
[PubMed:21166474] [WorldCat.org] [DOI] (I p) - ↑ 2.0 2.1 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
