Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_02098
- pan locus tag?: SAUPAN004899000
- symbol: SAOUHSC_02098
- pan gene symbol?: vraR
- synonym:
- product: DNA-binding response regulator VraR
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_02098
- symbol: SAOUHSC_02098
- product: DNA-binding response regulator VraR
- replicon: chromosome
- strand: -
- coordinates: 1972962..1973591
- length: 630
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3921170 NCBI
- RefSeq: YP_500589 NCBI
- BioCyc: G1I0R-1986 BioCyc
- MicrobesOnline: 1290545 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601ATGACGATTAAAGTATTGTTTGTGGATGATCATGAAATGGTACGTATAGGAATTTCAAGT
TATCTATCAACGCAAAGTGATATTGAAGTAGTTGGTGAAGGCGCTTCTGGTAAAGAAGCA
ATTGCCAAAGCCCATGAGTTGAAGCCAGATTTAATTTTAATGGATTTACTTATGGATGAC
ATGGATGGTGTAGAAGCGACGACTCAGATTAAAAAAGATTTACCGCAAATTAAAGTATTA
ATGTTAACTAGTTTTATTGAAGATAAAGAGGTATATCGTGCATTAGATGCAGGTGTCGAT
AGTTACATTTTAAAAACAACAAGTGCAAAAGATATCGCCGATGCAGTTCGTAAAACTTCT
AGAGGAGAATCTGTTTTTGAACCGGAAGTTTTAGTGAAAATGCGTAACCGTATGAAAAAG
CGCGCAGAGTTATATGAAATGCTTACAGAACGAGAAATGGAAATATTATTATTGATTGCG
AAAGGTTACTCAAATCAAGAAATTGCTAGTGCATCGCATATTACTATTAAAACGGTTAAG
ACACATGTGAGTAACATTTTAAGTAAGTTAGAAGTGCAAGATAGAACACAAGCTGTTATC
TATGCATTCCAACATAATTTAATTCAATAG60
120
180
240
300
360
420
480
540
600
630
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_02098
- symbol: SAOUHSC_02098
- description: DNA-binding response regulator VraR
- length: 209
- theoretical pI: 5.39084
- theoretical MW: 23545.2
- GRAVY: -0.121531
⊟Function[edit | edit source]
- TIGRFAM: Cellular processes Sporulation and germination sporulation transcription factor Spo0A (TIGR02875; HMM-score: 72.3)and 15 moretranscriptional regulator EpsA (TIGR03020; HMM-score: 55.8)Regulatory functions DNA interactions phosphate regulon transcriptional regulatory protein PhoB (TIGR02154; HMM-score: 55.6)Signal transduction Two-component systems phosphate regulon transcriptional regulatory protein PhoB (TIGR02154; HMM-score: 55.6)Regulatory functions DNA interactions heavy metal response regulator (TIGR01387; HMM-score: 36.5)Central intermediary metabolism Nitrogen metabolism nitrogen regulation protein NR(I) (TIGR01818; HMM-score: 36.5)Regulatory functions DNA interactions nitrogen regulation protein NR(I) (TIGR01818; HMM-score: 36.5)Signal transduction Two-component systems nitrogen regulation protein NR(I) (TIGR01818; HMM-score: 36.5)LuxR family transcriptional regulatory, chaperone HchA-associated (TIGR03541; HMM-score: 34)RNA polymerase sigma-70 factor, Bacteroides expansion family 1 (TIGR02985; HMM-score: 26.4)Signal transduction Two-component systems TMAO reductase sytem sensor TorS (TIGR02956; EC 2.7.13.3; HMM-score: 25.5)RNA polymerase sigma factor, sigma-70 family (TIGR02937; HMM-score: 21.1)Cellular processes Sporulation and germination RNA polymerase sigma-H factor (TIGR02859; HMM-score: 17.2)Transcription Transcription factors RNA polymerase sigma-H factor (TIGR02859; HMM-score: 17.2)proteobacterial dedicated sortase system response regulator (TIGR03787; HMM-score: 15.8)RNA polymerase sigma-70 factor, sigma-E family (TIGR02983; HMM-score: 13.4)
- TheSEED :
- Two component transcriptional regulator VraR
- PFAM: CheY (CL0304) Response_reg; Response regulator receiver domain (PF00072; HMM-score: 93.3)and 14 moreHTH (CL0123) GerE; Bacterial regulatory proteins, luxR family (PF00196; HMM-score: 71.1)Sigma70_r4_2; Sigma-70, region 4 (PF08281; HMM-score: 33.6)HTH_40; Helix-turn-helix domain (PF14493; HMM-score: 17.9)HTH_23; Homeodomain-like domain (PF13384; HMM-score: 16.6)UPF0122; Putative helix-turn-helix protein, YlxM / p13 like (PF04297; HMM-score: 15.6)Sigma70_r4; Sigma-70, region 4 (PF04545; HMM-score: 15.5)HTH_24; Winged helix-turn-helix DNA-binding (PF13412; HMM-score: 15.3)no clan defined DUF3194; Protein of unknown function (DUF3194) (PF11419; HMM-score: 15)PDDEXK (CL0236) PDDEXK_10; PD-(D/E)XK nuclease superfamily (PF07788; HMM-score: 14)HTH (CL0123) HTH_28; Helix-turn-helix domain (PF13518; HMM-score: 13.7)wHTH-PRTase_assc; wHTH-PRTase associated (PF24409; HMM-score: 13.6)no clan defined B12-binding; B12 binding domain (PF02310; HMM-score: 13)GT-A (CL0110) Rhamno_transf; Putative rhamnosyl transferase (PF11316; HMM-score: 12.9)HTH (CL0123) HTH_38; Helix-turn-helix domain (PF13936; HMM-score: 12.9)
⊟Structure, modifications & cofactors[edit | edit source]
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.97
- Cytoplasmic Membrane Score: 0
- Cellwall Score: 0.01
- Extracellular Score: 0.02
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.9867
- Cytoplasmic Membrane Score: 0.0016
- Cell wall & surface Score: 0.0001
- Extracellular Score: 0.0116
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.006867
- TAT(Tat/SPI): 0.000297
- LIPO(Sec/SPII): 0.000634
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MTIKVLFVDDHEMVRIGISSYLSTQSDIEVVGEGASGKEAIAKAHELKPDLILMDLLMDDMDGVEATTQIKKDLPQIKVLMLTSFIEDKEVYRALDAGVDSYILKTTSAKDIADAVRKTSRGESVFEPEVLVKMRNRMKKRAELYEMLTEREMEILLLIAKGYSNQEIASASHITIKTVKTHVSNILSKLEVQDRTQAVIYAFQHNLIQ
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas [6] [7]
- protein localization: data available for COL
- quantitative data / protein copy number per cell: data available for COL
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: SAOUHSC_02098 < SAOUHSC_02099 < SAOUHSC_02100 < SAOUHSC_02101predicted SigA promoter [8] : S823 < SAOUHSC_02098 < SAOUHSC_02099 < SAOUHSC_02100 < SAOUHSC_02101 < S824 < SAOUHSC_02102predicted SigA promoter [8] : SAOUHSC_02098 < SAOUHSC_02099 < SAOUHSC_02100 < SAOUHSC_02101 < S824 < SAOUHSC_02102 < SAOUHSC_A02013 < S826
⊟Regulation[edit | edit source]
- regulator: VraR* (activation) regulon
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [8]
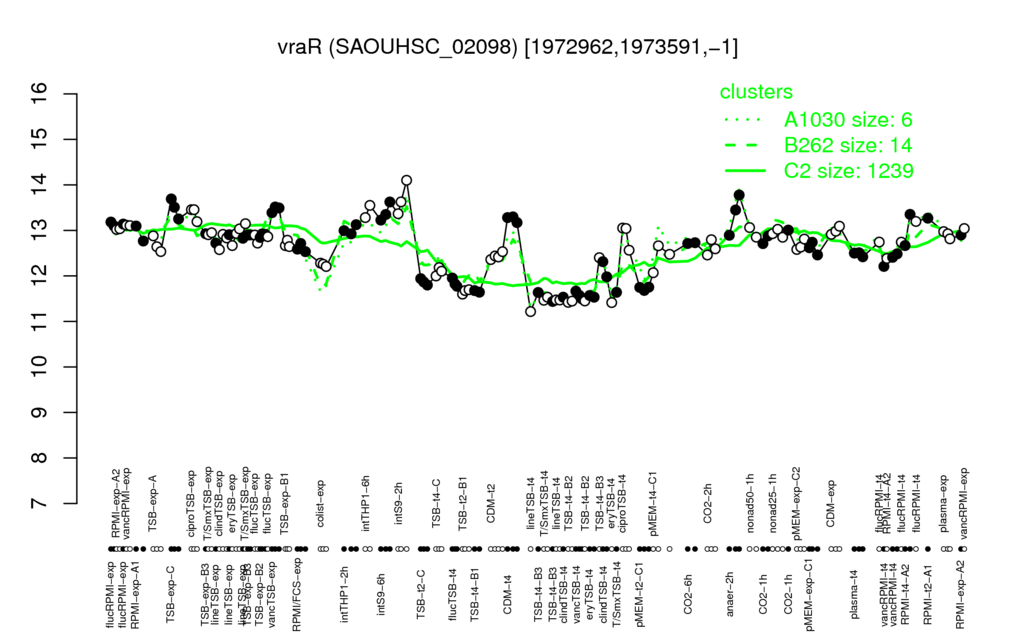 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Antoaneta Belcheva, Dasantila Golemi-Kotra
A close-up view of the VraSR two-component system. A mediator of Staphylococcus aureus response to cell wall damage.
J Biol Chem: 2008, 283(18);12354-64
[PubMed:18326495] [WorldCat.org] [DOI] (P p) - ↑ Pedro B Fernandes, Patricia Reed, João M Monteiro, Mariana G Pinho
Revisiting the Role of VraTSR in Staphylococcus aureus Response to Cell Wall-Targeting Antibiotics.
J Bacteriol: 2022, 204(8);e0016222
[PubMed:35862765] [WorldCat.org] [DOI] (I p) - ↑ Maëliss Germain, Hugo Robin, Kim Boi Le Huyen, Sébastien Massier, Nicolas Nalpas, Julie Hardouin, Philippe Bouloc, Astrid Rouillon, Svetlana Chabelskaya
sRNA-mediated crosstalk between cell wall stress and galactose metabolism in Staphylococcus aureus.
Nucleic Acids Res: 2025, 53(13);
[PubMed:40671522] [WorldCat.org] [DOI] (I p) - ↑ 4.0 4.1 Makoto Kuroda, Hiroko Kuroda, Taku Oshima, Fumihiko Takeuchi, Hirotada Mori, Keiichi Hiramatsu
Two-component system VraSR positively modulates the regulation of cell-wall biosynthesis pathway in Staphylococcus aureus.
Mol Microbiol: 2003, 49(3);807-21
[PubMed:12864861] [WorldCat.org] [DOI] (P p) - ↑ 5.0 5.1 Susan Boyle-Vavra, Shouhui Yin, Dae Sun Jo, Christopher P Montgomery, Robert S Daum
VraT/YvqF is required for methicillin resistance and activation of the VraSR regulon in Staphylococcus aureus.
Antimicrob Agents Chemother: 2013, 57(1);83-95
[PubMed:23070169] [WorldCat.org] [DOI] (I p) - ↑ Maren Depke, Stephan Michalik, Alexander Rabe, Kristin Surmann, Lars Brinkmann, Nico Jehmlich, Jörg Bernhardt, Michael Hecker, Bernd Wollscheid, Zhi Sun, Robert L Moritz, Uwe Völker, Frank Schmidt
A peptide resource for the analysis of Staphylococcus aureus in host-pathogen interaction studies.
Proteomics: 2015, 15(21);3648-61
[PubMed:26224020] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Maren Depke, Annette Murr, Manuela Gesell Salazar, Ulrike Kusebauch, Zhi Sun, Tanja C Meyer, Kristin Surmann, Henrike Pförtner, Petra Hildebrandt, Stefan Weiss, Laura Marcela Palma Medina, Melanie Gutjahr, Elke Hammer, Dörte Becher, Thomas Pribyl, Sven Hammerschmidt, Eric W Deutsch, Samuel L Bader, Michael Hecker, Robert L Moritz, Ulrike Mäder, Uwe Völker, Frank Schmidt
A global Staphylococcus aureus proteome resource applied to the in vivo characterization of host-pathogen interactions.
Sci Rep: 2017, 7(1);9718
[PubMed:28887440] [WorldCat.org] [DOI] (I e) - ↑ 8.0 8.1 8.2 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
