Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_02877
- pan locus tag?: SAUPAN006225000
- symbol: SAOUHSC_02877
- pan gene symbol?: crtN
- synonym:
- product: squalene synthase
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_02877
- symbol: SAOUHSC_02877
- product: squalene synthase
- replicon: chromosome
- strand: -
- coordinates: 2650767..2652275
- length: 1509
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3921548 NCBI
- RefSeq: YP_501332 NCBI
- BioCyc: G1I0R-2709 BioCyc
- MicrobesOnline: 1291303 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201
1261
1321
1381
1441
1501ATGAAGATTGCAGTAATTGGTGCAGGTGTCACAGGATTAGCAGCGGCAGCCCGTATTGCT
TCTCAAGGTCATGAAGTGACGATATTTGAAAAAAATAATAATGTAGGCGGGCGTATGAAT
CAATTAAAGAAAGACGGCTTTACATTTGATATGGGTCCCACAATTGTCATGATGCCAGAT
GTTTATAAAGATGTTTTTACAGCGTGTGGTAAAAATTATGAAGATTATATTGAATTGAGA
CAATTACGTTATATTTACGATGTGTATTTTGACCACGATGATCGTATAACGGTGCCTACA
GATTTAGCTGAATTACAGCAAATGCTAGAAAGTATAGAACCTGGTTCAACGCATGGTTTT
ATGTCCTTTTTAACGGATGTTTATAAAAAATATGAAATTGCACGTCGCTATTTCTTAGAA
AGAACGTATCGCAAACCGAGTGACTTTTATAATATGACGTCACTTGTGCAAGGTGCTAAG
TTAAAAACGTTAAATCATGCAGATCAGCTAATTGAACATTATATTGATAACGAAAAGATA
CAAAAGCTTTTAGCGTTTCAAACGTTATACATAGGAATTGATCCAAAACGAGGCCCGTCA
CTATATTCAATTATTCCTATGATTGAAATGATGTTTGGTGTGCATTTTATTAAAGGCGGT
ATGTATGGCATGGCTCAAGGGCTAGCGCAATTAAATAAAGACTTAGGCGTTAATATTGAA
CTAAATGCTGAAATTGAGCAAATTATTATTGATCCTAAATTCAAACGGGCCGATGCGATA
AAAGTGAATGGTGACATAAGAAAATTTGATAAAATTTTATGTACGGCTGATTTCCCTAGT
GTTGCGGAATCATTAATGCCAGATTTTGCACCTATTAAAAAGTATCCACCACATAAAATT
GCAGACTTAGATTACTCTTGTTCAGCATTTTTAATGTATATCGGTATAGATATTGATGTG
ACAGATCAAGTGAGACTTCATAATGTTATTTTTTCAGATGACTTTAGAGGCAATATTGAA
GAAATATTTGAGGGACGTTTATCATATGATCCTTCTATTTATGTGTATGTACCAGCGGTC
GCTGATAAATCACTTGCGCCAGAAGGCAAAACTGGTATTTATGTGCTAATGCCGACGCCG
GAACTTAAAACAGGTAGCGGAATCGATTGGTCAGATGAAGCTTTGACGCAACAAATAAAG
GAAATTATTTATCGTAAATTAGCAACGATTGAAGTATTTGAAGATATAAAATCGCATATT
GTTTCAGAAACAATCTTTACGCCAAATGATTTTGAGCAAACGTATCATGCGAAATTTGGT
TCGGCATTCGGTTTAATGCCAACTTTAGCGCAAAGTAATTATTATCGTCCACAAAATGTA
TCGCGAGATTATAAAGATTTATATTTTGCAGGTGCAAGTACGCATCCAGGTGCAGGCGTT
CCTATTGTCTTAACGAGTGCGAAAATAACTGTAGATGAAATGATTAAAGATATTGAGCGG
GGCGTATAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1260
1320
1380
1440
1500
1509
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_02877
- symbol: SAOUHSC_02877
- description: squalene synthase
- length: 502
- theoretical pI: 5.15038
- theoretical MW: 56741.7
- GRAVY: -0.17012
⊟Function[edit | edit source]
- reaction: EC 1.3.8.2? ExPASy4,4'-diapophytoene desaturase (4,4'-diapolycopene-forming) 15-cis-4,4'-diapophytoene + 4 FAD = all-trans-4,4'-diapolycopene + 4 FADH2
- TIGRFAM: Biosynthesis of cofactors, prosthetic groups, and carriers Other phytoene desaturase (TIGR02734; EC 1.14.99.-; HMM-score: 437.7)and 39 moreBiosynthesis of cofactors, prosthetic groups, and carriers Other C-3',4' desaturase CrtD (TIGR02733; EC 1.3.99.-; HMM-score: 79.5)Biosynthesis of cofactors, prosthetic groups, and carriers Other carotene isomerase (TIGR02730; EC 5.-.-.-; HMM-score: 75)Biosynthesis of cofactors, prosthetic groups, and carriers Heme, porphyrin, and cobalamin protoporphyrinogen oxidase (TIGR00562; EC 1.3.3.4; HMM-score: 47.5)squalene-associated FAD-dependent desaturase (TIGR03467; HMM-score: 43.9)mycofactocin system FadH/OYE family oxidoreductase 2 (TIGR03997; EC 1.-.-.-; HMM-score: 32.7)glutamate synthase, NADH/NADPH, small subunit (TIGR01317; EC 1.4.1.-; HMM-score: 31.6)9,9'-di-cis-zeta-carotene desaturase (TIGR02732; EC 1.3.5.6; HMM-score: 30)putative selenate reductase, YgfK subunit (TIGR03315; HMM-score: 27.9)Unknown function Enzymes of unknown specificity flavoprotein, HI0933 family (TIGR00275; HMM-score: 26.5)nucleotide sugar dehydrogenase (TIGR03026; HMM-score: 26.5)Amino acid biosynthesis Glutamate family glutamate synthase (NADPH), homotetrameric (TIGR01316; EC 1.4.1.13; HMM-score: 26)Cell envelope Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides UDP-galactopyranose mutase (TIGR00031; EC 5.4.99.9; HMM-score: 25.8)Protein synthesis tRNA and rRNA base modification tRNA U-34 5-methylaminomethyl-2-thiouridine biosynthesis protein MnmC, C-terminal domain (TIGR03197; HMM-score: 25.2)FAD dependent oxidoreductase TIGR03364 (TIGR03364; HMM-score: 24.8)Biosynthesis of cofactors, prosthetic groups, and carriers Thiamine glycine oxidase ThiO (TIGR02352; EC 1.4.3.19; HMM-score: 23.5)glutamate synthase, small subunit (TIGR01318; HMM-score: 21.8)Energy metabolism Amino acids and amines sarcosine oxidase, alpha subunit family (TIGR01372; HMM-score: 21)Biosynthesis of cofactors, prosthetic groups, and carriers Thiamine thiazole biosynthesis enzyme (TIGR00292; HMM-score: 18.8)Energy metabolism Electron transport flavocytochrome c (TIGR01813; HMM-score: 18.5)Biosynthesis of cofactors, prosthetic groups, and carriers Chlorophyll and bacteriochlorphyll geranylgeranyl reductase family (TIGR02032; EC 1.3.1.-; HMM-score: 17.8)Biosynthesis of cofactors, prosthetic groups, and carriers Other phytoene desaturase (TIGR02731; EC 1.14.99.-; HMM-score: 17.6)dihydrolipoyl dehydrogenase (TIGR01350; EC 1.8.1.4; HMM-score: 17.2)Biosynthesis of cofactors, prosthetic groups, and carriers Chlorophyll and bacteriochlorphyll geranylgeranyl reductase (TIGR02023; EC 1.3.1.-; HMM-score: 16.2)Biosynthesis of cofactors, prosthetic groups, and carriers Pantothenate and coenzyme A 2-dehydropantoate 2-reductase (TIGR00745; EC 1.1.1.-; HMM-score: 15)mycofactocin system FadH/OYE family oxidoreductase 1 (TIGR03996; EC 1.-.-.-; HMM-score: 14.1)Biosynthesis of cofactors, prosthetic groups, and carriers Pyridine nucleotides L-aspartate oxidase (TIGR00551; EC 1.4.3.16; HMM-score: 13.4)Energy metabolism Electron transport glutathione-disulfide reductase (TIGR01421; EC 1.8.1.7; HMM-score: 13.4)Unknown function Enzymes of unknown specificity putative bacillithiol system oxidoreductase, YpdA family (TIGR04018; EC 1.8.-.-; HMM-score: 13.2)Biosynthesis of cofactors, prosthetic groups, and carriers Menaquinone and ubiquinone ubiquinone biosynthesis hydroxylase, UbiH/UbiF/VisC/COQ6 family (TIGR01988; EC 1.14.13.-; HMM-score: 13)Protein synthesis tRNA and rRNA base modification tRNA:m(5)U-54 methyltransferase (TIGR00137; EC 2.1.1.74; HMM-score: 12.9)Biosynthesis of cofactors, prosthetic groups, and carriers Heme, porphyrin, and cobalamin precorrin 3B synthase CobZ (TIGR02485; HMM-score: 12.6)Energy metabolism Electron transport glutathione-disulfide reductase (TIGR01424; EC 1.8.1.7; HMM-score: 12.4)Energy metabolism Pentose phosphate pathway 6-phosphogluconate dehydrogenase (decarboxylating) (TIGR00872; EC 1.1.1.44; HMM-score: 11.8)Cell envelope Biosynthesis and degradation of murein sacculus and peptidoglycan UDP-N-acetylmuramoylalanine--D-glutamate ligase (TIGR01087; EC 6.3.2.9; HMM-score: 11.3)Cellular processes Detoxification CoA-disulfide reductase (TIGR03385; EC 1.8.1.14; HMM-score: 11.3)Energy metabolism Anaerobic glycerol-3-phosphate dehydrogenase, anaerobic, B subunit (TIGR03378; EC 1.1.5.3; HMM-score: 11)3-hydroxyacyl-CoA dehydrogenase PaaC (TIGR02279; EC 1.1.1.-; HMM-score: 10.9)Energy metabolism Electron transport NAD(P)(+) transhydrogenase (AB-specific), alpha subunit (TIGR00561; EC 1.6.1.2; HMM-score: 10.5)lycopene cyclase family protein (TIGR01790; HMM-score: 10.5)
- TheSEED :
- 4,4'-diapophytoene desaturase (4,4'-diapolycopene-forming) (EC 1.3.8.2)
- Dehydrosqualene desaturase (EC 1.3.8.2) CrtN
- PFAM: NADP_Rossmann (CL0063) Amino_oxidase; Flavin containing amine oxidoreductase (PF01593; HMM-score: 108.3)and 24 moreNAD_binding_8; NAD(P)-binding Rossmann-like domain (PF13450; HMM-score: 58)DAO; FAD dependent oxidoreductase (PF01266; HMM-score: 35)Pyr_redox_2; Pyridine nucleotide-disulphide oxidoreductase (PF07992; HMM-score: 28)HI0933_like; HI0933-like protein Rossmann domain (PF03486; HMM-score: 25.2)FAD_binding_2; FAD binding domain (PF00890; HMM-score: 24.9)Pyr_redox; Pyridine nucleotide-disulphide oxidoreductase (PF00070; HMM-score: 24.5)FAD_oxidored; FAD dependent oxidoreductase (PF12831; HMM-score: 24)AlaDh_PNT_C; Alanine dehydrogenase/PNT, C-terminal domain (PF01262; HMM-score: 21.4)UDPG_MGDP_dh_N; UDP-glucose/GDP-mannose dehydrogenase family, NAD binding domain (PF03721; HMM-score: 20.9)FMO-like; Flavin-binding monooxygenase-like (PF00743; HMM-score: 20.5)NAD_Gly3P_dh_N; NAD-dependent glycerol-3-phosphate dehydrogenase N-terminus (PF01210; HMM-score: 19.7)3HCDH_N; 3-hydroxyacyl-CoA dehydrogenase, NAD binding domain (PF02737; HMM-score: 19.6)Thi4; Thi4 family (PF01946; HMM-score: 18.2)MurF-HprK_N (CL0365) MurD-like_N; Mur ligase MurD-like, N-terminal domain (PF21799; HMM-score: 17.4)NADP_Rossmann (CL0063) GIDA; Glucose inhibited division protein A (PF01134; HMM-score: 17.2)NAD_binding_2; NAD binding domain of 6-phosphogluconate dehydrogenase (PF03446; HMM-score: 16.9)ApbA; Ketopantoate reductase PanE/ApbA (PF02558; HMM-score: 16.7)NAD_binding_9; FAD-NAD(P)-binding (PF13454; HMM-score: 16.5)MCRA; MCRA family (PF06100; HMM-score: 15.9)F420_oxidored; NADP oxidoreductase coenzyme F420-dependent (PF03807; HMM-score: 14.9)FAD_binding_3; FAD binding domain (PF01494; HMM-score: 13.9)TrkA_N; TrkA-N domain (PF02254; HMM-score: 13.7)KARI_N; Acetohydroxy acid isomeroreductase, NADPH-binding domain (PF07991; HMM-score: 12.4)Pyr_redox_3; Pyridine nucleotide-disulphide oxidoreductase (PF13738; HMM-score: 12.4)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors: FAD
- effectors:
⊟Localization[edit | edit source]
- PSORTb: unknown (no significant prediction)
- Cytoplasmic Score: 2.5
- Cytoplasmic Membrane Score: 2.5
- Cellwall Score: 2.5
- Extracellular Score: 2.5
- Internal Helices: 0
- DeepLocPro: Cytoplasmic Membrane
- Cytoplasmic Score: 0.177
- Cytoplasmic Membrane Score: 0.8022
- Cell wall & surface Score: 0.0007
- Extracellular Score: 0.0201
- LocateP: N-terminally anchored (No CS)
- Prediction by SwissProt Classification: Membrane
- Pathway Prediction: Sec-(SPI)
- Intracellular possibility: 0.5
- Signal peptide possibility: 0
- N-terminally Anchored Score: 7
- Predicted Cleavage Site: No CleavageSite
- SignalP: Signal peptide SP(Sec/SPI) length 23 aa
- SP(Sec/SPI): 0.653165
- TAT(Tat/SPI): 0.00485
- LIPO(Sec/SPII): 0.101583
- Cleavage Site: CS pos: 23-24. SQG-HE. Pr: 0.2989
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MKIAVIGAGVTGLAAAARIASQGHEVTIFEKNNNVGGRMNQLKKDGFTFDMGPTIVMMPDVYKDVFTACGKNYEDYIELRQLRYIYDVYFDHDDRITVPTDLAELQQMLESIEPGSTHGFMSFLTDVYKKYEIARRYFLERTYRKPSDFYNMTSLVQGAKLKTLNHADQLIEHYIDNEKIQKLLAFQTLYIGIDPKRGPSLYSIIPMIEMMFGVHFIKGGMYGMAQGLAQLNKDLGVNIELNAEIEQIIIDPKFKRADAIKVNGDIRKFDKILCTADFPSVAESLMPDFAPIKKYPPHKIADLDYSCSAFLMYIGIDIDVTDQVRLHNVIFSDDFRGNIEEIFEGRLSYDPSIYVYVPAVADKSLAPEGKTGIYVLMPTPELKTGSGIDWSDEALTQQIKEIIYRKLATIEVFEDIKSHIVSETIFTPNDFEQTYHAKFGSAFGLMPTLAQSNYYRPQNVSRDYKDLYFAGASTHPGAGVPIVLTSAKITVDEMIKDIERGV
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas [1] [2]
- protein localization: data available for COL
- quantitative data / protein copy number per cell:
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: SAOUHSC_02877 < SAOUHSC_02879 < SAOUHSC_02880 < SAOUHSC_02881 < SAOUHSC_02882predicted SigB promoter [3] : S1129 < SAOUHSC_02877 < SAOUHSC_02879 < SAOUHSC_02880 < SAOUHSC_02881 < SAOUHSC_02882 < S1130
⊟Regulation[edit | edit source]
- regulator: SigB* (activation) regulon
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [3]
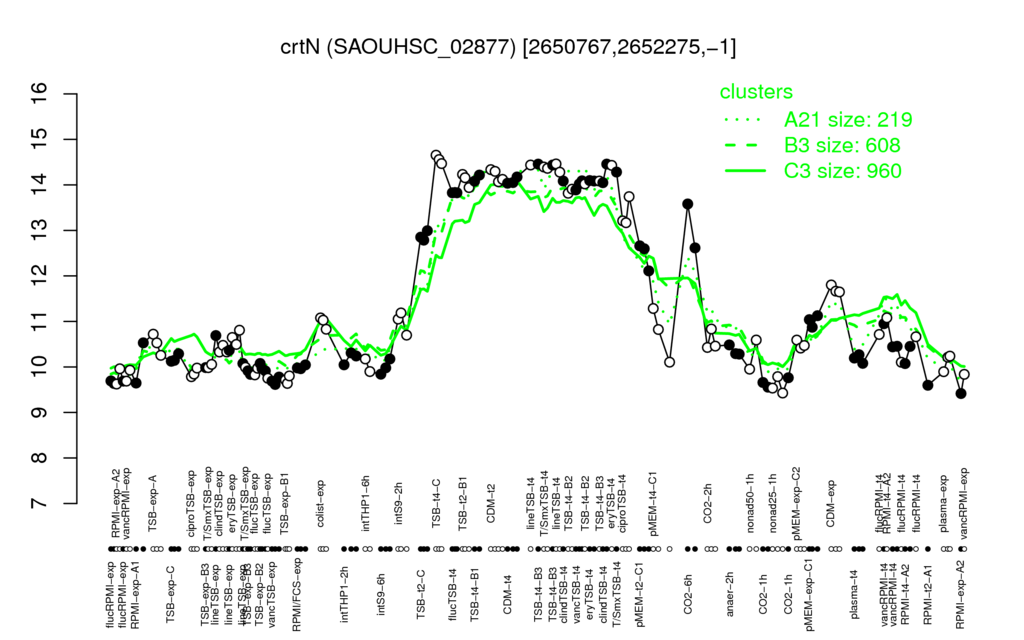 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Maren Depke, Stephan Michalik, Alexander Rabe, Kristin Surmann, Lars Brinkmann, Nico Jehmlich, Jörg Bernhardt, Michael Hecker, Bernd Wollscheid, Zhi Sun, Robert L Moritz, Uwe Völker, Frank Schmidt
A peptide resource for the analysis of Staphylococcus aureus in host-pathogen interaction studies.
Proteomics: 2015, 15(21);3648-61
[PubMed:26224020] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Maren Depke, Annette Murr, Manuela Gesell Salazar, Ulrike Kusebauch, Zhi Sun, Tanja C Meyer, Kristin Surmann, Henrike Pförtner, Petra Hildebrandt, Stefan Weiss, Laura Marcela Palma Medina, Melanie Gutjahr, Elke Hammer, Dörte Becher, Thomas Pribyl, Sven Hammerschmidt, Eric W Deutsch, Samuel L Bader, Michael Hecker, Robert L Moritz, Ulrike Mäder, Uwe Völker, Frank Schmidt
A global Staphylococcus aureus proteome resource applied to the in vivo characterization of host-pathogen interactions.
Sci Rep: 2017, 7(1);9718
[PubMed:28887440] [WorldCat.org] [DOI] (I e) - ↑ 3.0 3.1 3.2 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e) - ↑ Markus Bischoff, Paul Dunman, Jan Kormanec, Daphne Macapagal, Ellen Murphy, William Mounts, Brigitte Berger-Bächi, Steven Projan
Microarray-based analysis of the Staphylococcus aureus sigmaB regulon.
J Bacteriol: 2004, 186(13);4085-99
[PubMed:15205410] [WorldCat.org] [DOI] (P p)
