Jump to navigation
Jump to search
NCBI: 10-JUN-2013
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus COL
- locus tag: SACOL1637 [new locus tag: SACOL_RS08345 ]
- pan locus tag?: SAUPAN004163000
- symbol: dnaK
- pan gene symbol?: dnaK
- synonym:
- product: molecular chaperone DnaK
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SACOL1637 [new locus tag: SACOL_RS08345 ]
- symbol: dnaK
- product: molecular chaperone DnaK
- replicon: chromosome
- strand: -
- coordinates: 1666574..1668406
- length: 1833
- essential: unknown other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3238018 NCBI
- RefSeq: YP_186477 NCBI
- BioCyc: see SACOL_RS08345
- MicrobesOnline: 913086 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201
1261
1321
1381
1441
1501
1561
1621
1681
1741
1801ATGAGTAAAATTATTGGTATAGACTTAGGTACAACAAATTCATGTGTAACAGTATTAGAA
GGCGATGAGCCAAAAGTAATTCAAAACCCTGAAGGTTCACGTACAACACCATCTGTTGTA
GCTTTCAAAAATGGAGAAACTCAAGTTGGTGAAGTAGCAAAACGTCAAGCTATTACAAAC
CCAAACACTGTTCAATCTATTAAACGTCATATGGGTACTGATTATAAAGTAGATATTGAA
GGTAAATCATACACACCACAAGAAATCTCAGCTATGATTTTACAAAACTTAAAAAATACA
GCTGAAAGCTATTTAGGTGAGAAAGTTGACAAAGCTGTAATTACAGTACCTGCATACTTT
AACGATGCTGAACGTCAAGCAACTAAAGATGCTGGTAAAATTGCTGGTTTAGAAGTTGAG
CGTATCATTAATGAACCAACAGCTGCAGCATTAGCATATGGTTTAGACAAAACTGATAAA
GATGAAAAAGTTCTTGTTTTTGACTTAGGTGGCGGTACATTTGACGTATCTATCCTAGAA
TTAGGTGACGGTGTATTCGAAGTACTATCAACAGCCGGTGACAACAAACTTGGCGGTGAT
GATTTTGACCAAGTAATTATTGACTACCTAGTTGCAGAATTCAAAAAAGAAAATGGCGTA
GACTTATCTCAAGATAAAATGGCATTACAACGTTTGAAAGATGCTGCTGAAAAAGCTAAA
AAAGACTTATCAGGTGTATCACAAACTCAAATCTCATTACCATTTATCTCAGCTGGTGAA
AACGGTCCATTACACTTAGAAGTAAACTTAACTCGTTCTAAATTTGAAGAATTATCAGAT
TCATTAATTAGAAGAACAATGGAACCTACACGCCAAGCAATGAAAGACGCTGGCTTAACA
AACTCAGATATCGATGAAGTTATCTTAGTTGGTGGATCAACTCGTATTCCAGCAGTACAA
GAAGCTGTCAAAAAAGAAATCGGTAAAGAGCCTAACAAAGGAGTAAACCCGGACGAAGTA
GTGGCAATGGGAGCTGCAATCCAAGGTGGCGTTATCACAGGTGACGTTAAAGACGTAGTA
TTATTAGACGTAACACCACTATCTTTAGGTATTGAAATTTTAGGTGGACGTATGAATACG
TTAATTGAACGTAACACTACGATTCCTACATCTAAATCTCAAATCTATTCAACAGCAGTA
GATAATCAACCATCAGTAGATGTACACGTATTACAAGGTGAACGTCCAATGGCTGCGGAT
AATAAAACACTTGGTAGATTCCAATTGACTGATATTCCACCAGCTGAACGTGGTAAACCT
CAAATTGAAGTAACGTTTGATATCGATAAAAACGGTATTGTAAATGTAACTGCAAAAGAC
TTAGGTACAAATAAAGAACAAAGAATTACAATTCAATCAAGTTCTTCATTATCAGACGAA
GAAATCGACCGTATGGTAAAAGATGCTGAAGTTAACGCTGAAGCAGATAAAAAACGTCGT
GAAGAAGTAGACTTAAGAAACGAAGCTGACAGTCTAGTATTCCAAGTTGAAAAAACTTTA
ACTGATTTAGGCGAAAATATCGGTGAAGAAGATAAAAAATCTGCTGAAGAGAAAAAAGAC
GCTCTTAAAACTGCTTTAGAAGGTCAAGATATAGAAGATATTAAATCTAAAAAAGAAGAA
CTTGAAAAAGTGATTCAAGAATTATCAGCAAAAGTATATGAGCAAGCGGCTCAACAGCAA
CAACAAGCACAAGGTGCTAATGCTGGTCAAAACAATGATAGTACTGTAGAAGATGCTGAA
TTTAAAGAAGTAAAAGACGACGACAAAAAATAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1260
1320
1380
1440
1500
1560
1620
1680
1740
1800
1833
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SACOL1637 [new locus tag: SACOL_RS08345 ]
- symbol: DnaK
- description: molecular chaperone DnaK
- length: 610
- theoretical pI: 4.38499
- theoretical MW: 66360.9
- GRAVY: -0.523279
⊟Function[edit | edit source]
- TIGRFAM: Protein fate Protein folding and stabilization chaperone protein DnaK (TIGR02350; HMM-score: 938.7)and 4 moreProtein fate Protein folding and stabilization Fe-S protein assembly chaperone HscA (TIGR01991; HMM-score: 574.4)Cell envelope Biosynthesis and degradation of murein sacculus and peptidoglycan cell shape determining protein, MreB/Mrl family (TIGR00904; HMM-score: 66.9)ethanolamine utilization protein EutJ family protein (TIGR02529; HMM-score: 59.7)Central intermediary metabolism Phosphorus compounds exopolyphosphatase (TIGR03706; EC 3.6.1.11; HMM-score: 13)
- TheSEED :
- Chaperone protein DnaK
and 1 more - PFAM: Actin_ATPase (CL0108) HSP70; Hsp70 protein (PF00012; HMM-score: 834.9)and 3 moreMreB_Mbl; MreB/Mbl protein (PF06723; HMM-score: 71.9)ParM_N; Plasmid segregation protein ParM, C-terminal (PF21523; HMM-score: 17.9)Hydantoinase_A; Hydantoinase/oxoprolinase (PF01968; HMM-score: 17.8)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.97
- Cytoplasmic Membrane Score: 0
- Cellwall Score: 0.01
- Extracellular Score: 0.02
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.951
- Cytoplasmic Membrane Score: 0.0047
- Cell wall & surface Score: 0.0024
- Extracellular Score: 0.0419
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.01916
- TAT(Tat/SPI): 0.001154
- LIPO(Sec/SPII): 0.025443
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MSKIIGIDLGTTNSCVTVLEGDEPKVIQNPEGSRTTPSVVAFKNGETQVGEVAKRQAITNPNTVQSIKRHMGTDYKVDIEGKSYTPQEISAMILQNLKNTAESYLGEKVDKAVITVPAYFNDAERQATKDAGKIAGLEVERIINEPTAAALAYGLDKTDKDEKVLVFDLGGGTFDVSILELGDGVFEVLSTAGDNKLGGDDFDQVIIDYLVAEFKKENGVDLSQDKMALQRLKDAAEKAKKDLSGVSQTQISLPFISAGENGPLHLEVNLTRSKFEELSDSLIRRTMEPTRQAMKDAGLTNSDIDEVILVGGSTRIPAVQEAVKKEIGKEPNKGVNPDEVVAMGAAIQGGVITGDVKDVVLLDVTPLSLGIEILGGRMNTLIERNTTIPTSKSQIYSTAVDNQPSVDVHVLQGERPMAADNKTLGRFQLTDIPPAERGKPQIEVTFDIDKNGIVNVTAKDLGTNKEQRITIQSSSSLSDEEIDRMVKDAEVNAEADKKRREEVDLRNEADSLVFQVEKTLTDLGENIGEEDKKSAEEKKDALKTALEGQDIEDIKSKKEELEKVIQELSAKVYEQAAQQQQQAQGANAGQNNDSTVEDAEFKEVKDDDKK
⊟Experimental data[edit | edit source]
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
⊟Regulation[edit | edit source]
- regulator: HrcA (repression) regulon
HrcA (TF) important in Heat shock response; RegPrecise transcription unit transferred from N315 data RegPrecise
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: data available for NCTC8325
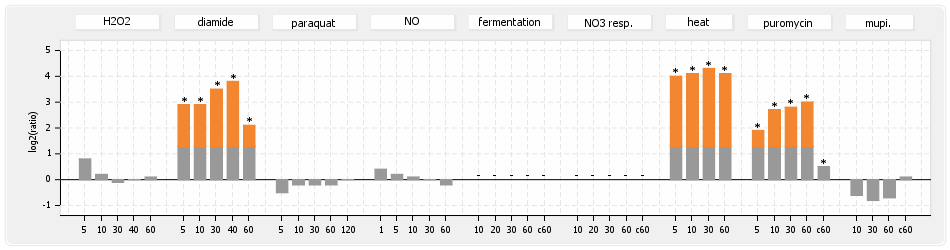
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: 32.19 h [8]
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Dörte Becher, Kristina Hempel, Susanne Sievers, Daniela Zühlke, Jan Pané-Farré, Andreas Otto, Stephan Fuchs, Dirk Albrecht, Jörg Bernhardt, Susanne Engelmann, Uwe Völker, Jan Maarten van Dijl, Michael Hecker
A proteomic view of an important human pathogen--towards the quantification of the entire Staphylococcus aureus proteome.
PLoS One: 2009, 4(12);e8176
[PubMed:19997597] [WorldCat.org] [DOI] (I e) - ↑ Kristina Hempel, Jan Pané-Farré, Andreas Otto, Susanne Sievers, Michael Hecker, Dörte Becher
Quantitative cell surface proteome profiling for SigB-dependent protein expression in the human pathogen Staphylococcus aureus via biotinylation approach.
J Proteome Res: 2010, 9(3);1579-90
[PubMed:20108986] [WorldCat.org] [DOI] (I p) - ↑ Kristina Hempel, Florian-Alexander Herbst, Martin Moche, Michael Hecker, Dörte Becher
Quantitative proteomic view on secreted, cell surface-associated, and cytoplasmic proteins of the methicillin-resistant human pathogen Staphylococcus aureus under iron-limited conditions.
J Proteome Res: 2011, 10(4);1657-66
[PubMed:21323324] [WorldCat.org] [DOI] (I p) - ↑ Andreas Otto, Jan Maarten van Dijl, Michael Hecker, Dörte Becher
The Staphylococcus aureus proteome.
Int J Med Microbiol: 2014, 304(2);110-20
[PubMed:24439828] [WorldCat.org] [DOI] (I p) - ↑ Daniela Zühlke, Kirsten Dörries, Jörg Bernhardt, Sandra Maaß, Jan Muntel, Volkmar Liebscher, Jan Pané-Farré, Katharina Riedel, Michael Lalk, Uwe Völker, Susanne Engelmann, Dörte Becher, Stephan Fuchs, Michael Hecker
Costs of life - Dynamics of the protein inventory of Staphylococcus aureus during anaerobiosis.
Sci Rep: 2016, 6;28172
[PubMed:27344979] [WorldCat.org] [DOI] (I e) - ↑ Artem Cherkasov, Michael Hsing, Roya Zoraghi, Leonard J Foster, Raymond H See, Nikolay Stoynov, Jihong Jiang, Sukhbir Kaur, Tian Lian, Linda Jackson, Huansheng Gong, Rick Swayze, Emily Amandoron, Farhad Hormozdiari, Phuong Dao, Cenk Sahinalp, Osvaldo Santos-Filho, Peter Axerio-Cilies, Kendall Byler, William R McMaster, Robert C Brunham, B Brett Finlay, Neil E Reiner
Mapping the protein interaction network in methicillin-resistant Staphylococcus aureus.
J Proteome Res: 2011, 10(3);1139-50
[PubMed:21166474] [WorldCat.org] [DOI] (I p) - ↑ Arnaud Chastanet, Juliette Fert, Tarek Msadek
Comparative genomics reveal novel heat shock regulatory mechanisms in Staphylococcus aureus and other Gram-positive bacteria.
Mol Microbiol: 2003, 47(4);1061-73
[PubMed:12581359] [WorldCat.org] [DOI] (P p) - ↑ Stephan Michalik, Jörg Bernhardt, Andreas Otto, Martin Moche, Dörte Becher, Hanna Meyer, Michael Lalk, Claudia Schurmann, Rabea Schlüter, Holger Kock, Ulf Gerth, Michael Hecker
Life and death of proteins: a case study of glucose-starved Staphylococcus aureus.
Mol Cell Proteomics: 2012, 11(9);558-70
[PubMed:22556279] [WorldCat.org] [DOI] (I p)
⊟Relevant publications[edit | edit source]
Vineet K Singh, Sugunya Utaida, Letitia S Jackson, R K Jayaswal, Brian J Wilkinson, Neal R Chamberlain
Role for dnaK locus in tolerance of multiple stresses in Staphylococcus aureus.
Microbiology (Reading): 2007, 153(Pt 9);3162-3173
[PubMed:17768259] [WorldCat.org] [DOI] (P p)