Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_00178
- pan locus tag?: SAUPAN001067000
- symbol: SAOUHSC_00178
- pan gene symbol?: —
- synonym:
- product: maltose ABC transporter permease
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_00178
- symbol: SAOUHSC_00178
- product: maltose ABC transporter permease
- replicon: chromosome
- strand: +
- coordinates: 195145..195984
- length: 840
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3919492 NCBI
- RefSeq: YP_498775 NCBI
- BioCyc: G1I0R-166 BioCyc
- MicrobesOnline: 1288669 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781ATGACAAAGAAGAAAAACATATTAAAAGCAATCGGTATTTACAGTTTTATAGCGATGATG
TTTGTCATCATTTTATATCCACTACTGTGGACATTTGGCATTTCCCTTAATCCAGGTACG
AACTTGTATGGTGCCAAAATGATACCAGACAATGCAACATTTAAAAATTATGCATTCTTA
CTATTCGATGACAGTAGTCAATACCTGACTTGGTATAAAAATACGCTTATCGTAGCATCT
GCAAATGCACTGTTTAGTGTGATATTTGTCACGTTAACAGCATATGCTTTTTCTAGATAT
CGCTTTGTTGGTCGTAAATACGGGCTGATTACATTTTTGATTTTACAAATGTTCCCTGTA
TTAATGGCAATGGTCGCAATCTATATTTTGCTAAATACAATTGGATTATTAGATTCTTTA
TTTGGACTAACACTGGTATATATTGGTGGATCAATACCGATGAATGCCTTTTTAGTGAAA
GGTTACTTCGATACGATTCCAAAAGAACTTGATGAATCTGCCAAAATTGATGGTGCAGGG
CATATGCGTATTTTCTTACAAATTATGCTTCCATTAGCTAAGCCGATTTTAGCAGTTGTT
GCTTTGTTCAATTTTATGGGGCCATTTATGGACTTTATATTACCTAAAATACTATTAAGA
AGTCCTGAAAAATTCACATTAGCAGTTGGATTGTTCAACTTTATTAATGATAAGTATGCA
AATAATTTCACAGTGTTTGCAGCAGGGGCAATTATGATTGCAGTACCTATAGCAATCGTA
TTCTTGTTCTTGCAACGCTATTTAGTATCAGGTTTAACAACAGGTGCGACAAAAGGTTAG60
120
180
240
300
360
420
480
540
600
660
720
780
840
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_00178
- symbol: SAOUHSC_00178
- description: maltose ABC transporter permease
- length: 279
- theoretical pI: 10.05
- theoretical MW: 31154.3
- GRAVY: 0.813978
⊟Function[edit | edit source]
- TIGRFAM: NifC-like ABC-type porter (TIGR01581; HMM-score: 29.1)Transport and binding proteins Amino acids, peptides and amines putative 2-aminoethylphosphonate ABC transporter, permease protein (TIGR03262; HMM-score: 28.1)Transport and binding proteins Anions sulfate ABC transporter, permease protein CysT (TIGR02139; HMM-score: 24.6)and 4 moreTransport and binding proteins Anions sulfate ABC transporter, permease protein (TIGR00969; HMM-score: 23.2)Transport and binding proteins Anions molybdate ABC transporter, permease protein (TIGR02141; HMM-score: 21.6)Transport and binding proteins Cations and iron carrying compounds K+-dependent Na+/Ca+ exchanger (TIGR00927; HMM-score: 8.3)Energy metabolism Electron transport cytochrome c oxidase, cbb3-type, CcoQ subunit (TIGR02736; EC 1.9.3.1; HMM-score: 6.6)
- TheSEED :
- Maltodextrin ABC transporter, permease protein MdxG
- PFAM: BPD_transp_1 (CL0404) BPD_transp_1; Binding-protein-dependent transport system inner membrane component (PF00528; HMM-score: 63.6)and 1 morePeptidase_MA (CL0126) DUF3267; Putative zincin peptidase (PF11667; HMM-score: 14.6)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic Membrane
- Cytoplasmic Score: 0
- Cytoplasmic Membrane Score: 10
- Cellwall Score: 0
- Extracellular Score: 0
- Internal Helices: 6
- DeepLocPro: Cytoplasmic Membrane
- Cytoplasmic Score: 0
- Cytoplasmic Membrane Score: 1
- Cell wall & surface Score: 0
- Extracellular Score: 0
- LocateP: Multi-transmembrane
- Prediction by SwissProt Classification: Membrane
- Pathway Prediction: Sec-(SPI)
- Intracellular possibility: 0.17
- Signal peptide possibility: -0.5
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.09566
- TAT(Tat/SPI): 0.001248
- LIPO(Sec/SPII): 0.006185
- predicted transmembrane helices (TMHMM): 6
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MTKKKNILKAIGIYSFIAMMFVIILYPLLWTFGISLNPGTNLYGAKMIPDNATFKNYAFLLFDDSSQYLTWYKNTLIVASANALFSVIFVTLTAYAFSRYRFVGRKYGLITFLILQMFPVLMAMVAIYILLNTIGLLDSLFGLTLVYIGGSIPMNAFLVKGYFDTIPKELDESAKIDGAGHMRIFLQIMLPLAKPILAVVALFNFMGPFMDFILPKILLRSPEKFTLAVGLFNFINDKYANNFTVFAAGAIMIAVPIAIVFLFLQRYLVSGLTTGATKG
⊟Experimental data[edit | edit source]
- experimentally validated:
- protein localization: data available for COL
- quantitative data / protein copy number per cell:
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
⊟Regulation[edit | edit source]
- regulators: MalR* (repression) regulon, CcpA* regulon
MalR* (TF) important in Maltose utilization, Maltodextrin utilization; RegPrecise transcription unit transferred from N315 data RegPrecise CcpA* (TF) important in Carbon catabolism; RegPrecise transcription unit transferred from N315 data RegPrecise
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [1]
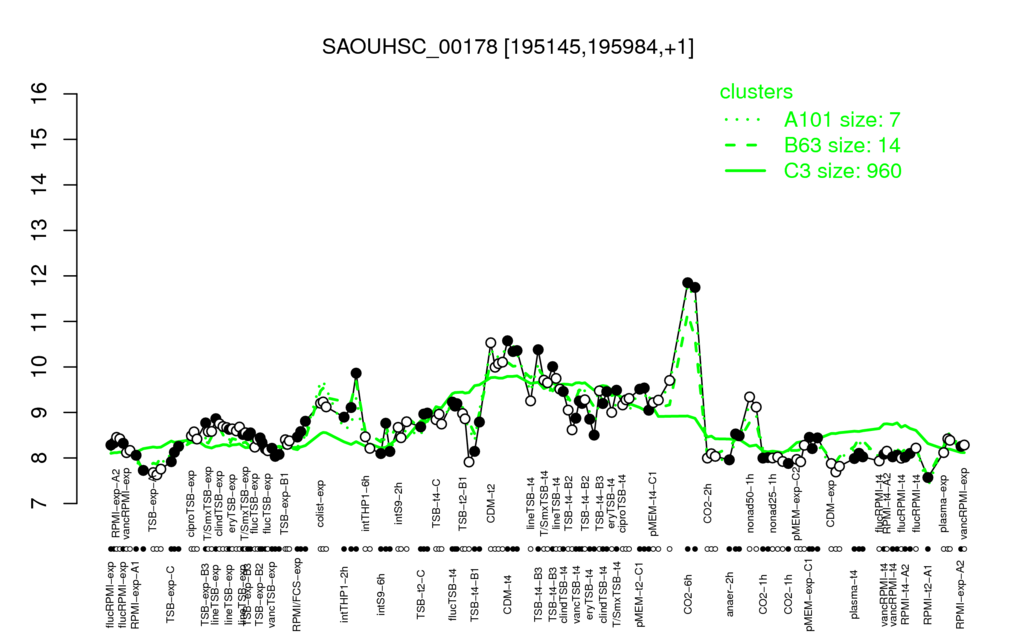 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
