Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_01724
- pan locus tag?: SAUPAN004218000
- symbol: SAOUHSC_01724
- pan gene symbol?: —
- synonym:
- product: hypothetical protein
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_01724
- symbol: SAOUHSC_01724
- product: hypothetical protein
- replicon: chromosome
- strand: -
- coordinates: 1630711..1631379
- length: 669
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3921074 NCBI
- RefSeq: YP_500233 NCBI
- BioCyc: G1I0R-1603 BioCyc
- MicrobesOnline: 1290147 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661ATGATAGATCAACAAACAATTTATCAATACATACAAAATGGAAAAATAGAAGAAGCGTTA
CAAGCATTGTTCGGAAATATCGAAGAAAATCCTACAATTATTGAAAATTATATTAATGCT
GGTATCGTACTTGCTGATGCGAATGAGATTGAAAAGGCAGAGCGTTTTTTCCAAAAAGCT
TTAACAATAGATCCGAAGAATGGCGTCGTATTTTTTAATCTAGCAAATGTATATTATAAT
CAGCAACGTTATCAAGAAGCTATTAAATTATATCAACAAGCATTACAAACAGAGATTGAA
CAAGTTGATTGTAATTATATGATCGGTATGGCGTTTAATCAGTTAGAATCATTTAAGCTG
GCATTGCCGTATTTAATGACTGCTGCGGAACTAGATAAAGACAAAGATGCAGAAGTTCAA
TTTCAATATGGTCTTGTATTATGTCAATTAGAAATGTTTAATGAAGCCATAACTCAACTT
AAACATGTATTAACGATTGATAAAAATCATGTTGATGCAAGATACAATTTGGGCTTAGCG
TTATTTATGAAAAATGAAGATATTGATGAAGCAATAACTCATTTTAAAGAAGCTGTGACT
ATCGACCCTAAACACTTATTAAGTCAGCATGCGCTGAAAACATTCACTAAAATGAAAGAG
GAGGAGTAA60
120
180
240
300
360
420
480
540
600
660
669
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_01724
- symbol: SAOUHSC_01724
- description: hypothetical protein
- length: 222
- theoretical pI: 4.41965
- theoretical MW: 25701.1
- GRAVY: -0.277477
⊟Function[edit | edit source]
- TIGRFAM: type IV pilus biogenesis/stability protein PilW (TIGR02521; HMM-score: 72.1)putative PEP-CTERM system TPR-repeat lipoprotein (TIGR02917; HMM-score: 70.8)and 8 moretype III secretion low calcium response chaperone LcrH/SycD (TIGR02552; HMM-score: 56.5)Transport and binding proteins Amino acids, peptides and amines mitochondrial precursor proteins import receptor (TIGR00990; HMM-score: 49.8)tol-pal system protein YbgF (TIGR02795; HMM-score: 31.4)putative peptide modification system cyclase (TIGR04510; HMM-score: 26.2)Unknown function General heme biosynthesis-associated TPR protein (TIGR00540; HMM-score: 24.5)pentatricopeptide repeat domain (TIGR00756; HMM-score: 21.9)Purines, pyrimidines, nucleosides, and nucleotides Salvage of nucleosides and nucleotides adenine phosphoribosyltransferase (TIGR01090; EC 2.4.2.7; HMM-score: 21.4)poly-beta-1,6 N-acetyl-D-glucosamine export porin PgaA (TIGR03939; HMM-score: 20.7)
- TheSEED :
- Tetratricopeptide repeat (TPR) family protein
- PFAM: TPR (CL0020) TPR_1; Tetratricopeptide repeat (PF00515; HMM-score: 99.7)TPR_2; Tetratricopeptide repeat (PF07719; HMM-score: 98.2)TPR_8; Tetratricopeptide repeat (PF13181; HMM-score: 93.3)TPR_11; TPR repeat (PF13414; HMM-score: 86.9)and 42 moreTPR_12; Tetratricopeptide repeat (PF13424; HMM-score: 78.9)TPR_19; Tetratricopeptide repeat (PF14559; HMM-score: 68.3)TPR_17; Tetratricopeptide repeat (PF13431; HMM-score: 66.1)TPR_14; Tetratricopeptide repeat (PF13428; HMM-score: 62.1)TPR_16; Tetratricopeptide repeat (PF13432; HMM-score: 61.8)ANAPC3; Anaphase-promoting complex, cyclosome, subunit 3 (PF12895; HMM-score: 56.2)TPR_10; Tetratricopeptide repeat (PF13374; HMM-score: 50.4)TPR_7; Tetratricopeptide repeat (PF13176; HMM-score: 47)TPR_CcmH_CycH; Cytochrome c-type biogenesis protein H TPR domain (PF23914; HMM-score: 45.1)TPR_6; Tetratricopeptide repeat (PF13174; HMM-score: 43.4)TPR_Slam; Surface lipoprotein assembly modifier, N-terminal TPR repeats (PF24575; HMM-score: 36.4)TPR_20; Tetratricopeptide repeat (PF14561; HMM-score: 34.2)T7SS_EccA1_N; T7SS, ESX-1 secretion system protein EccA1, N-terminal domain (PF21545; HMM-score: 32.4)TPR_9; Tetratricopeptide repeat (PF13371; HMM-score: 30.1)TPR-S; Tetratricopeptide Repeats-Sensor (PF20308; HMM-score: 29.5)TOM20_plant; Plant specific mitochondrial import receptor subunit TOM20 (PF06552; HMM-score: 28.4)ARM_TT21_C; Tetratricopeptide repeat protein 21 C-terminal ARM domain (PF25063; HMM-score: 26.7)PPR; PPR repeat (PF01535; HMM-score: 24.9)TPR_NPHP3; Nephrocystin-3 TPR domain (PF24885; HMM-score: 23.4)TPR_15; Tetratricopeptide repeat (PF13429; HMM-score: 21.9)ARM_TT21_4th; Tetratricopeptide repeat protein 21 forth ARM domain (PF25068; HMM-score: 21.8)MIT (CL0745) MIT; MIT (microtubule interacting and transport) domain (PF04212; HMM-score: 20.2)TPR (CL0020) TPR_3; Tetratricopeptide repeat (PF07720; HMM-score: 20.2)ARM_TT21_N; Tetratricopeptide repeat protein 21 N-terminal ARM repeat (PF25062; HMM-score: 20.2)ARM_TT21_5th; TT21 fifth ARM repeats domain (PF25064; HMM-score: 19.9)ARM_TT21; Tetratricopeptide repeat protein 21 ARM repeat (PF25058; HMM-score: 19.7)E_motif; E motif (PF20431; HMM-score: 18.1)NatA_aux_su; N-terminal acetyltransferase A, auxiliary subunit (PF12569; HMM-score: 17.4)PPR_2; PPR repeat family (PF13041; HMM-score: 17.3)no clan defined MatP; MatP N-terminal domain (PF06303; HMM-score: 16)TPR (CL0020) TPR_4; Tetratricopeptide repeat (PF07721; HMM-score: 15.8)TPR_P4H; Prolyl 4-hydroxylase peptide-substrate-binding domain (PF23558; HMM-score: 15.5)SRP_TPR_like; Putative TPR-like repeat (PF17004; HMM-score: 14.2)TPR_27; Plant tetratrico peptide repeats (PF23310; HMM-score: 14.2)no clan defined DUF1512; Protein of unknown function (DUF1512) N-terminal domain (PF07431; HMM-score: 13.6)TPR (CL0020) TPR_21; Tetratricopeptide repeat-like domain (PF09976; HMM-score: 12.8)TPR_MalT; MalT-like TPR region (PF17874; HMM-score: 12.8)Hect (CL0552) HECT_2; HECT-like Ubiquitin-conjugating enzyme (E2)-binding (PF09814; HMM-score: 12.2)TPR (CL0020) EAD11; Effector-associated domain 11 (PF19964; HMM-score: 12.1)Sel1; Sel1 repeat (PF08238; HMM-score: 11.7)Clathrin; Region in Clathrin and VPS (PF00637; HMM-score: 11.6)NADP_Rossmann (CL0063) Bin3; Bicoid-interacting protein 3 (Bin3) (PF06859; HMM-score: 9.3)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 7.5
- Cytoplasmic Membrane Score: 1.15
- Cellwall Score: 0.62
- Extracellular Score: 0.73
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.8886
- Cytoplasmic Membrane Score: 0.0614
- Cell wall & surface Score: 0.016
- Extracellular Score: 0.0341
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.004087
- TAT(Tat/SPI): 0.000163
- LIPO(Sec/SPII): 0.000383
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MIDQQTIYQYIQNGKIEEALQALFGNIEENPTIIENYINAGIVLADANEIEKAERFFQKALTIDPKNGVVFFNLANVYYNQQRYQEAIKLYQQALQTEIEQVDCNYMIGMAFNQLESFKLALPYLMTAAELDKDKDAEVQFQYGLVLCQLEMFNEAITQLKHVLTIDKNHVDARYNLGLALFMKNEDIDEAITHFKEAVTIDPKHLLSQHALKTFTKMKEEE
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas [1] [2]
- protein localization: data available for COL
- quantitative data / protein copy number per cell: data available for COL
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: SAOUHSC_01723 < SAOUHSC_01724predicted SigA promoter [3] : SAOUHSC_01719 < SAOUHSC_01720 < SAOUHSC_01721 < alaS < S677 < S678 < SAOUHSC_01723 < SAOUHSC_01724predicted SigA promoter [3] : SAOUHSC_01719 < SAOUHSC_01720 < SAOUHSC_01721 < alaS < S677 < S678 < SAOUHSC_01723 < SAOUHSC_01724 < S679 < SAOUHSC_01726 < SAOUHSC_01727 < S680
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [3]
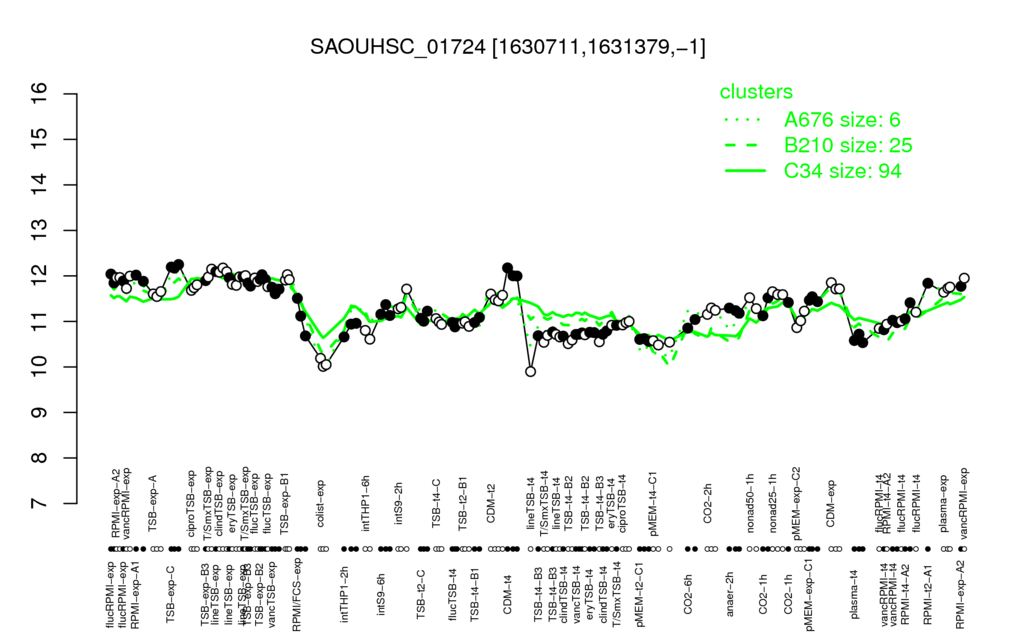 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Maren Depke, Stephan Michalik, Alexander Rabe, Kristin Surmann, Lars Brinkmann, Nico Jehmlich, Jörg Bernhardt, Michael Hecker, Bernd Wollscheid, Zhi Sun, Robert L Moritz, Uwe Völker, Frank Schmidt
A peptide resource for the analysis of Staphylococcus aureus in host-pathogen interaction studies.
Proteomics: 2015, 15(21);3648-61
[PubMed:26224020] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Maren Depke, Annette Murr, Manuela Gesell Salazar, Ulrike Kusebauch, Zhi Sun, Tanja C Meyer, Kristin Surmann, Henrike Pförtner, Petra Hildebrandt, Stefan Weiss, Laura Marcela Palma Medina, Melanie Gutjahr, Elke Hammer, Dörte Becher, Thomas Pribyl, Sven Hammerschmidt, Eric W Deutsch, Samuel L Bader, Michael Hecker, Robert L Moritz, Ulrike Mäder, Uwe Völker, Frank Schmidt
A global Staphylococcus aureus proteome resource applied to the in vivo characterization of host-pathogen interactions.
Sci Rep: 2017, 7(1);9718
[PubMed:28887440] [WorldCat.org] [DOI] (I e) - ↑ 3.0 3.1 3.2 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
