Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_00117
- pan locus tag?: SAUPAN000977000
- symbol: SAOUHSC_00117
- pan gene symbol?: capD
- synonym: cap8D
- product: capsular polysaccharide biosynthesis protein Cap5D
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_00117
- symbol: SAOUHSC_00117
- product: capsular polysaccharide biosynthesis protein Cap5D
- replicon: chromosome
- strand: +
- coordinates: 121649..123472
- length: 1824
- essential: no DEG other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3919826 NCBI
- RefSeq: YP_498717 NCBI
- BioCyc: G1I0R-108 BioCyc
- MicrobesOnline: 1288611 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201
1261
1321
1381
1441
1501
1561
1621
1681
1741
1801ATGGCACATTTATCTGTGAAATTGCGGCTTTTAATACTAGCATTAATCGATTCACTGATA
GTGACATTTTCAGTATTCGTAAGTTATTACATTTTAGAACCGTATTTCAAAACATATTCT
GTCAAATTATTAATATTGGCAGCTATATCACTATTCATATCGCATCATATTTCAGCATTT
ATTTTTAATATGTATCATCGAGCGTGGGAATATGCCAGTGTGAGTGAATTGATTTTAATT
GTTAAAGCTGTGACGACATCTATCGTTATTACGATGGTGGTCGTGACAATTGTTACAGGC
AATAGACCGTTTTTTAGATTGTATTTAATTACTTGGATGATGCACTTGATTTTAATAGGT
GGCTCAAGGTTATTTTGGCGTATTTATCGGAAATACCTTGGAGGTAAGTCATTTAATAAG
AAGCCAACTTTAGTTGTTGGTGCTGGTCAAGCAGGTTCAATGCTGATTAGACAAATGTTG
AAAAGTGACGAAATGAAACTTGAACCGGTATTAGCAGTCGATGATGACGAACATAAACGC
AATATCACAATTACTGAGGGTGTAAAAGTCCAAGGTAAAATTGCGGATATTCCAGAACTA
GTGAGGAAATATAAGATTAAAAAAATCATCATTGCAATTCCAACTATTGGTCAAGAGCGT
TTGAAAGAAATTAATAATATTTGCCATATGGATGGCGTTGAGTTATTGAAAATGCCAAAT
ATAGAAGACGTCATGTCTGGTGAGTTAGAAGTGAACCAACTTAAAAAAGTTGAAGTAGAA
GATTTACTAGGCAGAGATCCTGTTGAATTAGATATGGATATGATATCAAATGAATTGACG
AATAAAACTATTTTAGTTACGGGTGCAGGTGGTTCAATAGGATCAGAAATTTGTAGACAA
GTTTGTAATTTCTATCCAGAACGTATTATTCTACTTGGCCATGGTGAAAACAGTATTTAT
TTAATCAATCGTGAATTGCGAAATCGCTTCGGAAAAAATGTTGATATCGTTCCTATTATA
GCGGATGTGCAAAATAGAGCGCGTATGTTTGAAATTATGGAAACGTATAAACCATACGCA
GTTTATCATGCAGCAGCACACAAGCACGTGCCGTTAATGGAAGACAACCCTGAAGAAGCA
GTACGTAATAATATTTTAGGTACGAAAAATACTGCTGAAGCTGCTAAAAATGCAGAGGTA
AAGAAATTCGTTATGATTTCTACGGATAAAGCCGTTAATCCGCCTAATGTCATGGGAGCT
TCAAAGCGAATTGCAGAAATGATTATTCAAAGTTTAAATGATGAAACGCATCGAACAAAT
TTTGTTGCAGTGAGATTTGGTAATGTACTTGGATCGAGAGGATCTGTGATTCCACTTTTC
AAAAGTCAAATTGAAGAAGGTGGGCCAGTTACTGTGACACATCCTGAAATGACACGTTAC
TTTATGACAATTCCTGAAGCTTCTAGACTAGTTTTGCAGGCAGGGGCATTAGCAGAAGGT
GGCGAAGTATTTGTGCTAGATATGGGAGAACCAGTGAAAATTGTAGATTTGGCACGTAAT
TTAATTAAGCTAAGTGGTAAAAAAGAAGACGACATACGCATTACTTATACAGGGATTAGA
CCCGGCGAAAAAATGTTTGAAGAGCTTATGAATAAAGATGAGGTTCATCCTGAACAAGTA
TTTGAAAAAATTTATCGTGGCAAAGTACAACATATGAAATGTAATGAAGTTGAAGCGATT
ATTCAAGACATCGTCAATGACTTTAGTAAAGAAAAAATTATTAACTATGCCAATGGCAAA
AAGGGAGATAATTATGTTCGATGA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1260
1320
1380
1440
1500
1560
1620
1680
1740
1800
1824
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_00117
- symbol: SAOUHSC_00117
- description: capsular polysaccharide biosynthesis protein Cap5D
- length: 607
- theoretical pI: 8.49807
- theoretical MW: 68639.9
- GRAVY: -0.0014827
⊟Function[edit | edit source]
- TIGRFAM: UDP-N-acetylglucosamine 4,6-dehydratase (inverting) (TIGR03589; EC 4.2.1.115; HMM-score: 204.4)UDP-N-acetylglucosamine 4,6-dehydratase/5-epimerase (TIGR04130; EC 4.2.1.-,5.1.3.-; HMM-score: 164.9)and 15 moreexopolysaccharide biosynthesis polyprenyl glycosylphosphotransferase (TIGR03025; HMM-score: 98.1)undecaprenyl-phosphate glucose phosphotransferase (TIGR03023; EC 2.7.8.-; HMM-score: 87.4)Energy metabolism Sugars UDP-glucose 4-epimerase GalE (TIGR01179; EC 5.1.3.2; HMM-score: 63.2)Cell envelope Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides dTDP-glucose 4,6-dehydratase (TIGR01181; EC 4.2.1.46; HMM-score: 55)undecaprenyl-phosphate galactose phosphotransferase WbaP (TIGR03022; EC 2.7.8.6; HMM-score: 52.3)Cell envelope Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides dTDP-4-dehydrorhamnose reductase (TIGR01214; EC 1.1.1.133; HMM-score: 45.1)NAD dependent epimerase/dehydratase, LLPSF_EDH_00030 family (TIGR04180; HMM-score: 42.3)Cell envelope Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides CDP-glucose 4,6-dehydratase (TIGR02622; EC 4.2.1.45; HMM-score: 39.2)hopanoid-associated sugar epimerase (TIGR03466; HMM-score: 33.6)sugar transferase, PEP-CTERM system associated (TIGR03013; HMM-score: 29.9)Cell envelope Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides ADP-glyceromanno-heptose 6-epimerase (TIGR02197; EC 5.1.3.20; HMM-score: 24.4)rhamnulose-1-phosphate aldolase/alcohol dehydrogenase (TIGR02632; EC 1.1.1.1,4.1.2.19; HMM-score: 15.7)thioester reductase domain (TIGR01746; HMM-score: 15.2)Cell envelope Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides GDP-mannose 4,6-dehydratase (TIGR01472; EC 4.2.1.47; HMM-score: 14.5)2-hydroxycyclohexanecarboxyl-CoA dehydrogenase (TIGR03206; EC 1.1.1.-; HMM-score: 12.7)
- TheSEED :
- Capsular polysaccharide synthesis enzyme Cap8D
- PFAM: NADP_Rossmann (CL0063) Polysacc_synt_2; Polysaccharide biosynthesis protein (PF02719; HMM-score: 427.1)and 18 moreEpimerase; NAD dependent epimerase/dehydratase family (PF01370; HMM-score: 88.3)CoA_binding_3; CoA-binding domain (PF13727; HMM-score: 65.2)GDP_Man_Dehyd; GDP-mannose 4,6 dehydratase (PF16363; HMM-score: 57.3)RmlD_sub_bind; RmlD substrate binding domain (PF04321; HMM-score: 50.9)3Beta_HSD; 3-beta hydroxysteroid dehydrogenase/isomerase family (PF01073; HMM-score: 43.8)NAD_binding_10; NAD(P)H-binding (PF13460; HMM-score: 30.1)KR; KR domain (PF08659; HMM-score: 25.3)NAD_binding_4; Male sterility protein (PF07993; HMM-score: 22.3)adh_short; short chain dehydrogenase (PF00106; HMM-score: 20.5)GFO_IDH_MocA; Oxidoreductase family, NAD-binding Rossmann fold (PF01408; HMM-score: 17.4)TrkA_N; TrkA-N domain (PF02254; HMM-score: 16.6)NmrA; NmrA-like family (PF05368; HMM-score: 15.6)Shikimate_DH; Shikimate / quinate 5-dehydrogenase (PF01488; HMM-score: 14.2)no clan defined Tmemb_170; Putative transmembrane protein 170 (PF10190; HMM-score: 14.1)DUF3842; Domain of unknown function (DUF3842) (PF12953; HMM-score: 13.9)HTH (CL0123) wHTH_TTC3; TTC3-like, winged helix turn helix domain (PF24812; HMM-score: 13.3)NADP_Rossmann (CL0063) DapB_N; Dihydrodipicolinate reductase, N-terminus (PF01113; HMM-score: 13)DXP_reductoisom; 1-deoxy-D-xylulose 5-phosphate reductoisomerase (PF02670; HMM-score: 11)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic Membrane
- Cytoplasmic Score: 0
- Cytoplasmic Membrane Score: 10
- Cellwall Score: 0
- Extracellular Score: 0
- Internal Helices: 4
- DeepLocPro: Cytoplasmic Membrane
- Cytoplasmic Score: 0.0005
- Cytoplasmic Membrane Score: 0.998
- Cell wall & surface Score: 0
- Extracellular Score: 0.0014
- LocateP: Multi-transmembrane
- Prediction by SwissProt Classification: Membrane
- Pathway Prediction: Sec-(SPI)
- Intracellular possibility: 0.17
- Signal peptide possibility: -0.5
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.001647
- TAT(Tat/SPI): 0.000105
- LIPO(Sec/SPII): 0.008968
- predicted transmembrane helices (TMHMM): 4
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MAHLSVKLRLLILALIDSLIVTFSVFVSYYILEPYFKTYSVKLLILAAISLFISHHISAFIFNMYHRAWEYASVSELILIVKAVTTSIVITMVVVTIVTGNRPFFRLYLITWMMHLILIGGSRLFWRIYRKYLGGKSFNKKPTLVVGAGQAGSMLIRQMLKSDEMKLEPVLAVDDDEHKRNITITEGVKVQGKIADIPELVRKYKIKKIIIAIPTIGQERLKEINNICHMDGVELLKMPNIEDVMSGELEVNQLKKVEVEDLLGRDPVELDMDMISNELTNKTILVTGAGGSIGSEICRQVCNFYPERIILLGHGENSIYLINRELRNRFGKNVDIVPIIADVQNRARMFEIMETYKPYAVYHAAAHKHVPLMEDNPEEAVRNNILGTKNTAEAAKNAEVKKFVMISTDKAVNPPNVMGASKRIAEMIIQSLNDETHRTNFVAVRFGNVLGSRGSVIPLFKSQIEEGGPVTVTHPEMTRYFMTIPEASRLVLQAGALAEGGEVFVLDMGEPVKIVDLARNLIKLSGKKEDDIRITYTGIRPGEKMFEELMNKDEVHPEQVFEKIYRGKVQHMKCNEVEAIIQDIVNDFSKEKIINYANGKKGDNYVR
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas [1] [2]
- protein localization: data available for COL
- quantitative data / protein copy number per cell:
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- predicted SigA promoter [3] : S37 > SAOUHSC_00114 > SAOUHSC_00115 > SAOUHSC_00116 > SAOUHSC_00117 > SAOUHSC_00118 > SAOUHSC_00119 > SAOUHSC_00120 > SAOUHSC_00121 > SAOUHSC_00122 > SAOUHSC_00123 > SAOUHSC_00124 > SAOUHSC_00125 > SAOUHSC_00126 > SAOUHSC_00127predicted SigB promoter [3] : SAOUHSC_00114 > SAOUHSC_00115 > SAOUHSC_00116 > SAOUHSC_00117 > SAOUHSC_00118 > SAOUHSC_00119 > SAOUHSC_00120 > SAOUHSC_00121 > SAOUHSC_00122 > SAOUHSC_00123 > SAOUHSC_00124 > SAOUHSC_00125 > SAOUHSC_00126 > SAOUHSC_00127 > SAOUHSC_00128 > S38 > SAOUHSC_00129
⊟Regulation[edit | edit source]
- regulators: CodY* (repression) regulon, SigB* (activation) regulon
CodY* (TF) important in Amino acid metabolism; RegPrecise transcription unit transferred from N315 data RegPrecise SigB* (sigma factor) controlling a large regulon involved in stress/starvation response and adaptation; [6] [3] other strains
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [3]
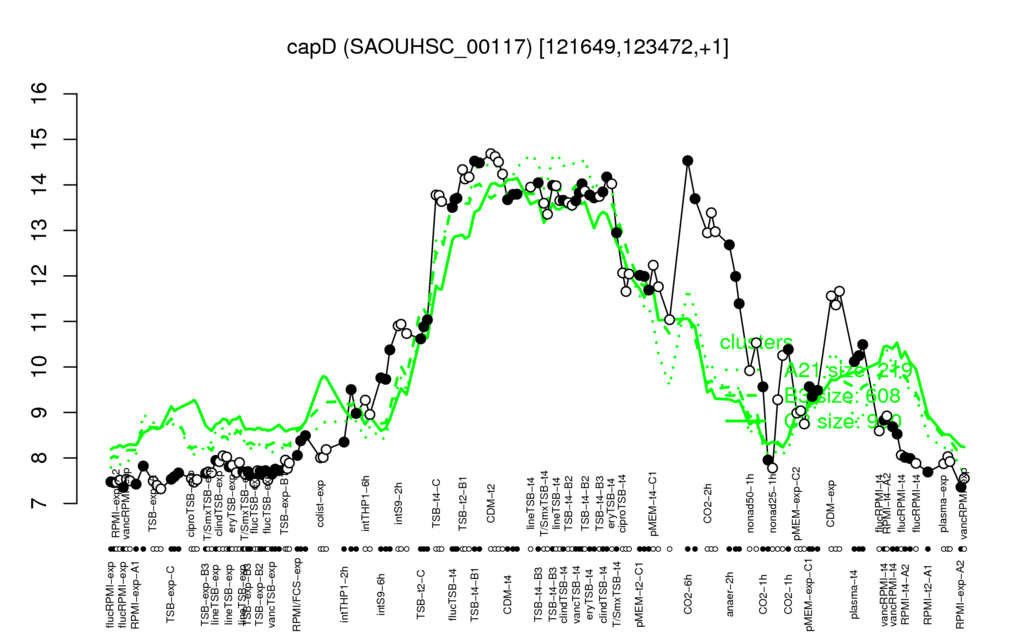 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Maren Depke, Stephan Michalik, Alexander Rabe, Kristin Surmann, Lars Brinkmann, Nico Jehmlich, Jörg Bernhardt, Michael Hecker, Bernd Wollscheid, Zhi Sun, Robert L Moritz, Uwe Völker, Frank Schmidt
A peptide resource for the analysis of Staphylococcus aureus in host-pathogen interaction studies.
Proteomics: 2015, 15(21);3648-61
[PubMed:26224020] [WorldCat.org] [DOI] (I p) - ↑ Stephan Michalik, Maren Depke, Annette Murr, Manuela Gesell Salazar, Ulrike Kusebauch, Zhi Sun, Tanja C Meyer, Kristin Surmann, Henrike Pförtner, Petra Hildebrandt, Stefan Weiss, Laura Marcela Palma Medina, Melanie Gutjahr, Elke Hammer, Dörte Becher, Thomas Pribyl, Sven Hammerschmidt, Eric W Deutsch, Samuel L Bader, Michael Hecker, Robert L Moritz, Ulrike Mäder, Uwe Völker, Frank Schmidt
A global Staphylococcus aureus proteome resource applied to the in vivo characterization of host-pathogen interactions.
Sci Rep: 2017, 7(1);9718
[PubMed:28887440] [WorldCat.org] [DOI] (I e) - ↑ 3.0 3.1 3.2 3.3 Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e) - ↑ Daniela Keinhörster, Shilpa Elizabeth George, Christopher Weidenmaier, Christiane Wolz
Function and regulation of Staphylococcus aureus wall teichoic acids and capsular polysaccharides.
Int J Med Microbiol: 2019, 309(6);151333
[PubMed:31362856] [WorldCat.org] [DOI] (I p) - ↑ S Sau, J Sun, C Y Lee
Molecular characterization and transcriptional analysis of type 8 capsule genes in Staphylococcus aureus.
J Bacteriol: 1997, 179(5);1614-21
[PubMed:9045821] [WorldCat.org] [DOI] (P p) - ↑ Markus Bischoff, Paul Dunman, Jan Kormanec, Daphne Macapagal, Ellen Murphy, William Mounts, Brigitte Berger-Bächi, Steven Projan
Microarray-based analysis of the Staphylococcus aureus sigmaB regulon.
J Bacteriol: 2004, 186(13);4085-99
[PubMed:15205410] [WorldCat.org] [DOI] (P p)
