Jump to navigation
Jump to search
NCBI: 03-AUG-2016
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus NCTC8325
- locus tag: SAOUHSC_03008
- pan locus tag?: SAUPAN006428000
- symbol: SAOUHSC_03008
- pan gene symbol?: hisF
- synonym:
- product: multifunctional imidazole glycerol phosphate synthase subunit/phosphoribosyl-AMP cyclohydrolase/phosphoribosyl-ATP pyrophosphatase
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SAOUHSC_03008
- symbol: SAOUHSC_03008
- product: multifunctional imidazole glycerol phosphate synthase subunit/phosphoribosyl-AMP cyclohydrolase/phosphoribosyl-ATP pyrophosphatase
- replicon: chromosome
- strand: -
- coordinates: 2782800..2783559
- length: 759
- essential: no DEG other strains
- comment: The sequence of SAOUHSC_03008 was corrected based on the resequencing performed by Berscheid et al., 2012 [1], resulting in a shorter CDS.
⊟Accession numbers[edit | edit source]
- Gene ID: 3921489 NCBI
- RefSeq: YP_501457 NCBI
- BioCyc: G1I0R-2829 BioCyc
- MicrobesOnline: 1291428 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721ATGATTAAAAAACGTATCATTCCATGTTTAGATGTCAAAGATGGTCGTGTCGTTAAAGGG
ATTCAATTTAAAGGATTAAGGGATATTGGGAATCCTGTTGATTTAGCAATGTATTACAAT
GAAGCGGGTGCTGATGAATTAGTATTTTTAGACATCTCTAAGACGGAAGAGGGTCATAGC
TTAATGCTAGAAGTGATTGAACAGACAGCGTCACGCTTGTTTATCCCTCTTACTGTAGGG
GGTGGGATTCAAAGTCTCGATGATATTACCCAATTGCTAAATCATGGTGCAGATAAAGTA
TCATTAAATTCAAGTGCTTTAAAAAATCCACAGCTCATTAAACAAGCGAGTGATAAATTC
GGTAGACAATGCATCTGCATAGCAATTGATAGCTATTATGATCCTGAAAGAAAAGCACAT
TATTGTTGTACGACTGGTGGTAAAAAAATGACAAATATTAAAGTATATGACTGGGTACAG
CAAGTAGAACAGTTAGGTGCAGGTGAGCTCCTCGTTACAAGTATGGGACATGATGGTATG
AAACAAGGCTTTGATATTGAACACCTAGCAAATATTAAGTCTCTTGTAAATATTCCAATC
ATTGCTTCTGGTGGTGGTGGCAATGCACAACACTTTGTAGAATTATTTGATCAGACGGAT
GTTTCTGCAGGTTTAGCTGCAAGTATATTACATGATCGAGAAACGACGGTTCAATCTATT
AAAGAAGTGATACGGCAAGGGGGTATAGCAGTAAGATGA60
120
180
240
300
360
420
480
540
600
660
720
759
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SAOUHSC_03008
- symbol: SAOUHSC_03008
- description: multifunctional imidazole glycerol phosphate synthase subunit/phosphoribosyl-AMP cyclohydrolase/phosphoribosyl-ATP pyrophosphatase
- length: 252
- theoretical pI: 6.15747
- theoretical MW: 27498.4
- GRAVY: -0.097619
⊟Function[edit | edit source]
- reaction: 5-[(5-phospho-1-deoxyribulos-1-ylamino)methylideneamino]-1-(5-phosphoribosyl)imidazole-4-carboxamide + L-glutamine = imidazole-glycerol phosphate + 5-aminoimidazol-4-carboxamide ribonucleotide + L-glutamate + H2O?
- TIGRFAM: Amino acid biosynthesis Histidine family imidazoleglycerol phosphate synthase, cyclase subunit (TIGR00735; HMM-score: 334.3)glycosyl amidation-associated protein WbuZ (TIGR03572; HMM-score: 271.3)and 12 moreAmino acid biosynthesis Histidine family 1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]imidazole-4-carboxamide isomerase (TIGR00007; EC 5.3.1.16; HMM-score: 118.7)bifunctional HisA/TrpF protein (TIGR01919; EC 5.3.1.16,5.3.1.24; HMM-score: 74.2)Unknown function General hisA/hisF family protein (TIGR00734; HMM-score: 34.6)Amino acid biosynthesis Histidine family phosphoribosylformimino-5-aminoimidazole carboxamide ribotide isomerase (TIGR02129; EC 5.3.1.16; HMM-score: 33.2)Biosynthesis of cofactors, prosthetic groups, and carriers Thiamine thiamine-phosphate diphosphorylase (TIGR00693; EC 2.5.1.3; HMM-score: 24.3)Unknown function General putative TIM-barrel protein, nifR3 family (TIGR00737; HMM-score: 18.7)geranylgeranylglyceryl phosphate synthase family protein (TIGR01768; EC 2.5.1.-; HMM-score: 16.2)dihydroorotate dehydrogenase family protein (TIGR01037; EC 1.3.-.-; HMM-score: 14.3)Energy metabolism Glycolysis/gluconeogenesis fructose-1,6-bisphosphate aldolase, class II (TIGR01859; EC 4.1.2.13; HMM-score: 14.2)Protein synthesis tRNA and rRNA base modification tRNA dihydrouridine synthase A (TIGR00742; EC 1.-.-.-; HMM-score: 13.6)mycofactocin system FadH/OYE family oxidoreductase 2 (TIGR03997; EC 1.-.-.-; HMM-score: 12.9)phosphoglycerol geranylgeranyltransferase (TIGR01769; EC 2.5.1.41; HMM-score: 12.3)
- TheSEED :
- Imidazole glycerol phosphate synthase cyclase subunit (EC 4.1.3.-)
- Phosphoribosyl-AMP cyclohydrolase (EC 3.5.4.19)
- Phosphoribosyl-ATP pyrophosphatase (EC 3.6.1.31)
Amino Acids and Derivatives Histidine Metabolism Histidine Biosynthesis Imidazole glycerol phosphate synthase cyclase subunit (EC 4.1.3.-)and 5 moreAmino Acids and Derivatives Histidine Metabolism Histidine Biosynthesis Phosphoribosyl-AMP cyclohydrolase (EC 3.5.4.19)Amino Acids and Derivatives Histidine Metabolism Histidine Biosynthesis Phosphoribosyl-ATP pyrophosphatase (EC 3.6.1.31)Amino Acids and Derivatives Histidine Metabolism Histidine Biosynthesis in plants Imidazole glycerol phosphate synthase cyclase subunit (EC 4.1.3.-) - PFAM: TIM_barrel (CL0036) His_biosynth; Histidine biosynthesis protein (PF00977; HMM-score: 260.5)and 6 moreDus; Dihydrouridine synthase (Dus) (PF01207; HMM-score: 32.3)TMP-TENI; Thiamine monophosphate synthase (PF02581; HMM-score: 27.3)NMO; Nitronate monooxygenase (PF03060; HMM-score: 22.1)G3P_antiterm; Glycerol-3-phosphate responsive antiterminator (PF04309; HMM-score: 20.3)DHO_dh; Dihydroorotate dehydrogenase (PF01180; HMM-score: 13.1)ThiG; Thiazole biosynthesis protein ThiG (PF05690; HMM-score: 12.5)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.67
- Cytoplasmic Membrane Score: 0.01
- Cellwall Score: 0.15
- Extracellular Score: 0.17
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.9582
- Cytoplasmic Membrane Score: 0.0171
- Cell wall & surface Score: 0.0002
- Extracellular Score: 0.0244
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: -1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.003209
- TAT(Tat/SPI): 0.000081
- LIPO(Sec/SPII): 0.000378
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MIKKRIIPCLDVKDGRVVKGIQFKGLRDIGNPVDLAMYYNEAGADELVFLDISKTEEGHSLMLEVIEQTASRLFIPLTVGGGIQSLDDITQLLNHGADKVSLNSSALKNPQLIKQASDKFGRQCICIAIDSYYDPERKAHYCCTTGGKKMTNIKVYDWVQQVEQLGAGELLVTSMGHDGMKQGFDIEHLANIKSLVNIPIIASGGGGNAQHFVELFDQTDVSAGLAASILHDRETTVQSIKEVIRQGGIAVR
⊟Experimental data[edit | edit source]
- experimentally validated:
- protein localization: data available for COL
- quantitative data / protein copy number per cell:
- interaction partners:
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: SAOUHSC_03008 < SAOUHSC_03009 < hisH < hisB < SAOUHSC_03012 < SAOUHSC_03013 < hisG < hisZ
⊟Regulation[edit | edit source]
- regulators: CodY* (repression) regulon, HisR* (repression) regulon
CodY* (TF) important in Amino acid metabolism; RegPrecise transcription unit transferred from N315 data RegPrecise HisR* (TF) important in Histidine biosynthesis; RegPrecise transcription unit transferred from N315 data RegPrecise
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: [2]
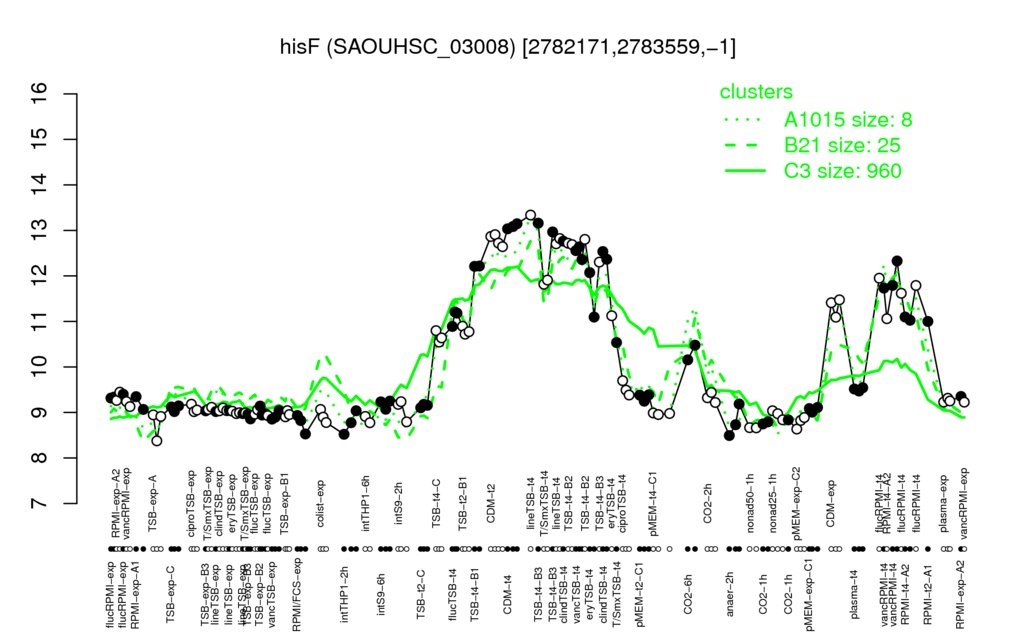 Multi-gene expression profiles
Multi-gene expression profiles
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
- Aureolib: no data available
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Anne Berscheid, Peter Sass, Konstantin Weber-Lassalle, Ambrose L Cheung, Gabriele Bierbaum
Revisiting the genomes of the Staphylococcus aureus strains NCTC 8325 and RN4220.
Int J Med Microbiol: 2012, 302(2);84-7
[PubMed:22417616] [WorldCat.org] [DOI] (I p) - ↑ Ulrike Mäder, Pierre Nicolas, Maren Depke, Jan Pané-Farré, Michel Debarbouille, Magdalena M van der Kooi-Pol, Cyprien Guérin, Sandra Dérozier, Aurelia Hiron, Hanne Jarmer, Aurélie Leduc, Stephan Michalik, Ewoud Reilman, Marc Schaffer, Frank Schmidt, Philippe Bessières, Philippe Noirot, Michael Hecker, Tarek Msadek, Uwe Völker, Jan Maarten van Dijl
Staphylococcus aureus Transcriptome Architecture: From Laboratory to Infection-Mimicking Conditions.
PLoS Genet: 2016, 12(4);e1005962
[PubMed:27035918] [WorldCat.org] [DOI] (I e)
