Jump to navigation
Jump to search
NCBI: 10-JUN-2013
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus COL
- locus tag: SACOL0816 [new locus tag: SACOL_RS04195 ]
- pan locus tag?: SAUPAN002662000
- symbol: secA
- pan gene symbol?: secA
- synonym:
- product: preprotein translocase subunit SecA
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SACOL0816 [new locus tag: SACOL_RS04195 ]
- symbol: secA
- product: preprotein translocase subunit SecA
- replicon: chromosome
- strand: +
- coordinates: 839016..841547
- length: 2532
- essential: unknown other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3236839 NCBI
- RefSeq: YP_185690 NCBI
- BioCyc: see SACOL_RS04195
- MicrobesOnline: 912288 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201
1261
1321
1381
1441
1501
1561
1621
1681
1741
1801
1861
1921
1981
2041
2101
2161
2221
2281
2341
2401
2461
2521ATGGGATTTTTATCAAAAATTCTTGATGGCAATAATAAAGAAATTAAACAGTTAGGTAAA
CTTGCTGATAAAGTAATCGCTTTAGAAGAAAAAACGGCAATTTTAACTGATGAAGAAATT
CGTAATAAAACGAAACAATTCCAAACAGAATTAGCTGACATTGATAATGTCAAAAAGCAA
AATGATTATTTAGATAAAATTTTACCAGAAGCATATGCACTTGTTAGAGAAGGCTCTAAA
CGTGTATTCAATATGACACCATATAAAGTTCAAATTATGGGTGGTATTGCAATTCATAAA
GGTGATATCGCTGAGATGAGAACAGGTGAAGGTAAAACATTAACAGCGACAATGCCAACA
TACTTAAATGCATTAGCTGGTAGAGGTGTTCACGTTATTACAGTCAATGAATACTTATCA
AGTGTTCAAAGTGAAGAAATGGCTGAGTTATATAACTTCTTAGGTTTGACTGTCGGATTA
AACTTAAACAGTAAGACGACAGAAGAAAAACGTGAAGCATACGCACAAGACATTACTTAC
AGTACTAATAATGAGCTAGGTTTTGATTACTTACGAGATAACATGGTGAATTATTCTGAA
GATAGAGTAATGCGTCCATTACATTTTGCAATCATTGATGAGGTTGACTCAATTTTAATC
GACGAGGCACGTACGCCATTAATTATTTCTGGTGAAGCTGAAAAGTCAACGTCACTTTAT
ACACAAGCAAATGTTTTTGCGAAAATGTTAAAACAGGACGAAGATTATAAATACGATGAA
AAAACGAAAGCTGTACATTTAACAGAACAAGGTGCGGATAAAGCTGAACGTATGTTCAAA
GTTGAAAACTTATATGATGTACAAAATGTTGATGTTATTAGTCATATCAACACAGCTTTA
CGTGCGCACGTTACATTACAACGTGACGTAGACTATATGGTTGTTGATGGCGAAGTATTA
ATTGTCGATCAATTTACAGGACGTACAATGCCAGGCCGTCGTTTCTCGGAAGGTTTACAC
CAAGCTATTGAAGCGAAGGAAGGCGTTCAAATTCAAAATGAATCTAAAACTATGGCGTCT
ATTACATTCCAAAACTATTTCAGAATGTACAATAAACTTGCGGGTATGACAGGTACAGCT
AAAACTGAAGAAGAAGAATTTAGAAATATTTATAACATGACAGTAACTCAAATTCCGACA
AATAAACCTGTGCAACGTAACGATAAGTCTGATTTAATTTACATTAGCCAAAAAGGTAAA
TTTGATGCAGTAGTAGAAGATGTTGTTGAAAAACACAAGGCAGGGCAACCAGTGCTATTA
GGTACTGTTGCAGTTGAGACTTCTGAATATATTTCAAATTTACTTAAAAAACGTGGTATC
CGTCATGATGTGTTAAATGCGAAAAATCATGAACGTGAAGCTGAAATTGTTGCAGGCGCT
GGACAAAAAGGTGCCGTTACTATTGCCACTAACATGGCTGGTCGTGGTACAGATATCAAA
TTAGGTGAAGGCGTAGAGGAATTAGGCGGTTTAGCAGTAATAGGTACAGAGCGACATGAA
TCTCGTCGTATTGATGACCAGTTACGTGGTCGTTCTGGACGTCAAGGTGATAAAGGGGAT
AGTCGCTTCTATTTATCATTACAAGATGAATTAATGATTCGTTTTGGTTCTGAACGTTTA
CAGAAAATGATGAGCCGACTAGGTTTAGATGACTCTACACCAATTGAATCAAAAATGGTA
TCAAGAGCTGTAGAATCAGCACAAAAACGTGTAGAAGGTAATAACTTCGACGCGCGTAAA
CGTATCTTAGAATACGATGAAGTATTACGTAAACAACGTGAAATTATCTATAACGAAAGA
AATAGTATTATTGATGAAGAAGACAGCTCTCAAGTTGTAGATGCAATGCTACGTTCAACG
TTACAACGTAGTATCAATTACTATATTAATACAGCAGATGACGAGCCTGAATATCAACCA
TTCATCGACTACATTAATGACATCTTCTTACAAGAAGGTGACATTACAGAGGATGATATC
AAAGGTAAAGATGCTGAAGATATTTTCGAAGTCGTTTGGGCTAAGATTGAAGCAGCATAT
CAAAGTCAAAAAGATATCTTAGAAGAACAAATGAATGAGTTTGAGCGTATGATTTTACTT
CGTTCTATTGATAGCCATTGGACTGATCATATCGACACAATGGATCAATTACGTCAAGGT
ATTCACTTACGTTCTTATGCACAACAAAATCCATTACGTGACTATCAAAATGAAGGTCAT
GAATTATTTGATATCATGATGCAAAATATTGAAGAAGATACTTGTAAATTCATTTTAAAA
TCTGTAGTACAAGTTGAAGATAATATTGAACGTGAAAAAACAACAGAGTTTGGTGAAGCG
AAGCACGTTTCAGCTGAAGATGGTAAAGAAAAAGTGAAACCGAAACCAATCGTTAAAGGC
GATCAAGTTGGTCGTAACGATGATTGTCCATGTGGTAGTGGTAAAAAATTCAAAAATTGC
CATGGAAAATAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1260
1320
1380
1440
1500
1560
1620
1680
1740
1800
1860
1920
1980
2040
2100
2160
2220
2280
2340
2400
2460
2520
2532
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SACOL0816 [new locus tag: SACOL_RS04195 ]
- symbol: SecA
- description: preprotein translocase subunit SecA
- length: 843
- theoretical pI: 4.86612
- theoretical MW: 95959.5
- GRAVY: -0.62242
⊟Function[edit | edit source]
- TIGRFAM: Protein fate Protein and peptide secretion and trafficking preprotein translocase, SecA subunit (TIGR00963; HMM-score: 1129)Protein fate Protein and peptide secretion and trafficking accessory Sec system translocase SecA2 (TIGR04397; HMM-score: 933.2)and 3 moreProtein fate Protein and peptide secretion and trafficking accessory Sec system translocase SecA2 (TIGR03714; HMM-score: 852)accessory Sec system translocase SecA2 (TIGR04221; HMM-score: 615.2)SWIM/SEC-C metal-binding motif protein, PBPRA1643 family (TIGR04102; HMM-score: 25)
- TheSEED :
- Protein translocase subunit SecA
and 1 more - PFAM: P-loop_NTPase (CL0023) SecA_DEAD; SecA DEAD-like domain (PF07517; HMM-score: 401.4)and 8 moreP-loop_SecA; SecA P-loop domain (PF21090; HMM-score: 257.8)no clan defined SecA_SW; SecA Wing and Scaffold domain (PF07516; HMM-score: 207.9)P-loop_NTPase (CL0023) SecA_PP_bind; SecA preprotein cross-linking domain (PF01043; HMM-score: 146.7)C4-like (CL0821) SEC-C; SEC-C motif (PF02810; HMM-score: 34.1)P-loop_NTPase (CL0023) DEAD; DEAD/DEAH box helicase (PF00270; HMM-score: 29.8)no clan defined DUF6429; Domain of unknown function (DUF6429) (PF20008; HMM-score: 17.2)P-loop_NTPase (CL0023) Helicase_C; Helicase conserved C-terminal domain (PF00271; HMM-score: 17.1)no clan defined DUF5663; Protein of unknown function (DUF5663) (PF18908; HMM-score: 10.4)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors: Zn2+
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.97
- Cytoplasmic Membrane Score: 0
- Cellwall Score: 0.01
- Extracellular Score: 0.02
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.4865
- Cytoplasmic Membrane Score: 0.4847
- Cell wall & surface Score: 0.0034
- Extracellular Score: 0.0254
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.001789
- TAT(Tat/SPI): 0.000231
- LIPO(Sec/SPII): 0.000414
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MGFLSKILDGNNKEIKQLGKLADKVIALEEKTAILTDEEIRNKTKQFQTELADIDNVKKQNDYLDKILPEAYALVREGSKRVFNMTPYKVQIMGGIAIHKGDIAEMRTGEGKTLTATMPTYLNALAGRGVHVITVNEYLSSVQSEEMAELYNFLGLTVGLNLNSKTTEEKREAYAQDITYSTNNELGFDYLRDNMVNYSEDRVMRPLHFAIIDEVDSILIDEARTPLIISGEAEKSTSLYTQANVFAKMLKQDEDYKYDEKTKAVHLTEQGADKAERMFKVENLYDVQNVDVISHINTALRAHVTLQRDVDYMVVDGEVLIVDQFTGRTMPGRRFSEGLHQAIEAKEGVQIQNESKTMASITFQNYFRMYNKLAGMTGTAKTEEEEFRNIYNMTVTQIPTNKPVQRNDKSDLIYISQKGKFDAVVEDVVEKHKAGQPVLLGTVAVETSEYISNLLKKRGIRHDVLNAKNHEREAEIVAGAGQKGAVTIATNMAGRGTDIKLGEGVEELGGLAVIGTERHESRRIDDQLRGRSGRQGDKGDSRFYLSLQDELMIRFGSERLQKMMSRLGLDDSTPIESKMVSRAVESAQKRVEGNNFDARKRILEYDEVLRKQREIIYNERNSIIDEEDSSQVVDAMLRSTLQRSINYYINTADDEPEYQPFIDYINDIFLQEGDITEDDIKGKDAEDIFEVVWAKIEAAYQSQKDILEEQMNEFERMILLRSIDSHWTDHIDTMDQLRQGIHLRSYAQQNPLRDYQNEGHELFDIMMQNIEEDTCKFILKSVVQVEDNIEREKTTEFGEAKHVSAEDGKEKVKPKPIVKGDQVGRNDDCPCGSGKKFKNCHGK
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas
- protein localization: Cytoplasmic [1] [2] [3] [4]
- quantitative data / protein copy number per cell: 380 [5]
- interaction partners:
SACOL1199 (ftsZ) cell division protein FtsZ [6] (data from MRSA252) SACOL0593 (fusA) elongation factor G [6] (data from MRSA252) SACOL1513 (hup) DNA-binding protein HU [6] (data from MRSA252) SACOL0533 (metS) methionyl-tRNA synthetase [6] (data from MRSA252) SACOL1102 (pdhA) pyruvate dehydrogenase complex E1 component subunit alpha [6] (data from MRSA252) SACOL1104 (pdhC) branched-chain alpha-keto acid dehydrogenase E2 [6] (data from MRSA252) SACOL1105 (pdhD) dihydrolipoamide dehydrogenase [6] (data from MRSA252) SACOL1745 (pyk) pyruvate kinase [6] (data from MRSA252) SACOL2238 (rplD) 50S ribosomal protein L4 [6] (data from MRSA252) SACOL1257 (rplS) 50S ribosomal protein L19 [6] (data from MRSA252) SACOL1274 (rpsB) 30S ribosomal protein S2 [6] (data from MRSA252) SACOL2230 (rpsQ) 30S ribosomal protein S17 [6] (data from MRSA252) SACOL0594 (tuf) elongation factor Tu [6] (data from MRSA252) SACOL0944 NADH dehydrogenase [6] (data from MRSA252) SACOL0973 fumarylacetoacetate hydrolase [6] (data from MRSA252)
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: no polycistronic organisation predicted
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: data available for NCTC8325
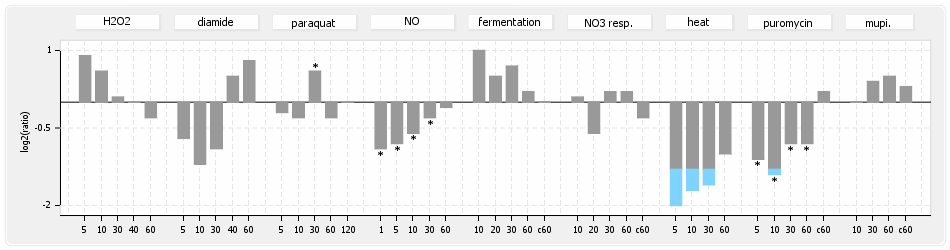
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Dörte Becher, Kristina Hempel, Susanne Sievers, Daniela Zühlke, Jan Pané-Farré, Andreas Otto, Stephan Fuchs, Dirk Albrecht, Jörg Bernhardt, Susanne Engelmann, Uwe Völker, Jan Maarten van Dijl, Michael Hecker
A proteomic view of an important human pathogen--towards the quantification of the entire Staphylococcus aureus proteome.
PLoS One: 2009, 4(12);e8176
[PubMed:19997597] [WorldCat.org] [DOI] (I e) - ↑ Kristina Hempel, Jan Pané-Farré, Andreas Otto, Susanne Sievers, Michael Hecker, Dörte Becher
Quantitative cell surface proteome profiling for SigB-dependent protein expression in the human pathogen Staphylococcus aureus via biotinylation approach.
J Proteome Res: 2010, 9(3);1579-90
[PubMed:20108986] [WorldCat.org] [DOI] (I p) - ↑ Kristina Hempel, Florian-Alexander Herbst, Martin Moche, Michael Hecker, Dörte Becher
Quantitative proteomic view on secreted, cell surface-associated, and cytoplasmic proteins of the methicillin-resistant human pathogen Staphylococcus aureus under iron-limited conditions.
J Proteome Res: 2011, 10(4);1657-66
[PubMed:21323324] [WorldCat.org] [DOI] (I p) - ↑ Andreas Otto, Jan Maarten van Dijl, Michael Hecker, Dörte Becher
The Staphylococcus aureus proteome.
Int J Med Microbiol: 2014, 304(2);110-20
[PubMed:24439828] [WorldCat.org] [DOI] (I p) - ↑ Daniela Zühlke, Kirsten Dörries, Jörg Bernhardt, Sandra Maaß, Jan Muntel, Volkmar Liebscher, Jan Pané-Farré, Katharina Riedel, Michael Lalk, Uwe Völker, Susanne Engelmann, Dörte Becher, Stephan Fuchs, Michael Hecker
Costs of life - Dynamics of the protein inventory of Staphylococcus aureus during anaerobiosis.
Sci Rep: 2016, 6;28172
[PubMed:27344979] [WorldCat.org] [DOI] (I e) - ↑ 6.00 6.01 6.02 6.03 6.04 6.05 6.06 6.07 6.08 6.09 6.10 6.11 6.12 6.13 6.14 Artem Cherkasov, Michael Hsing, Roya Zoraghi, Leonard J Foster, Raymond H See, Nikolay Stoynov, Jihong Jiang, Sukhbir Kaur, Tian Lian, Linda Jackson, Huansheng Gong, Rick Swayze, Emily Amandoron, Farhad Hormozdiari, Phuong Dao, Cenk Sahinalp, Osvaldo Santos-Filho, Peter Axerio-Cilies, Kendall Byler, William R McMaster, Robert C Brunham, B Brett Finlay, Neil E Reiner
Mapping the protein interaction network in methicillin-resistant Staphylococcus aureus.
J Proteome Res: 2011, 10(3);1139-50
[PubMed:21166474] [WorldCat.org] [DOI] (I p)