Jump to navigation
Jump to search
NCBI: 10-JUN-2013
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus COL
- locus tag: SACOL1385 [new locus tag: SACOL_RS07050 ]
- pan locus tag?: SAUPAN003735000
- symbol: acnA
- pan gene symbol?: citB
- synonym:
- product: aconitate hydratase
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SACOL1385 [new locus tag: SACOL_RS07050 ]
- symbol: acnA
- product: aconitate hydratase
- replicon: chromosome
- strand: +
- coordinates: 1390934..1393639
- length: 2706
- essential: unknown other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3238094 NCBI
- RefSeq: YP_186238 NCBI
- BioCyc: see SACOL_RS07050
- MicrobesOnline: 912844 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721
781
841
901
961
1021
1081
1141
1201
1261
1321
1381
1441
1501
1561
1621
1681
1741
1801
1861
1921
1981
2041
2101
2161
2221
2281
2341
2401
2461
2521
2581
2641
2701ATGGCTGCAAATTTTAAAGAGCAATCAAAAAAACATTTTGACTTGAATGGCCAAAGTTAT
ACTTACTATGATTTAAAAGCTGTAGAAGAGCAAGGTATTACTAAAGTTTCCAATTTACCT
TATTCAATTCGTGTTTTGTTAGAATCTTTACTTCGTCAAGAAGATGATTTTGTAATTACA
GACGATCATATTAAAGCTTTAAGTCAGTTTGGAAAAGATGGAAATGAAGGCGAGGTACCA
TTTAAACCTTCTCGTGTTATTTTACAAGATTTCACAGGTGTACCAGCCGTAGTTGATTTA
GCTTCTTTACGTAAAGCAATGGATGACGTTGGGGGAGATATTACTAAAATTAATCCAGAA
GTACCGGTGGATTTAGTTATTGACCACTCAGTTCAAGTGGATAGCTATGCAAATCCAGAA
GCTCTTGAACGTAATATGAAATTAGAATTTGAACGTAACTATGAACGTTATCAGTTTTTA
AATTGGGCAACGAAAGCATTTGATAATTACAATGCAGTTCCTCCTGCAACTGGAATAGTT
CACCAAGTTAACTTAGAATATTTAGCAAGTGTTGTACATGTTCGTGATGTAGATGGTGAA
AAAACTGCATTTCCAGATACATTAGTTGGTACTGATTCACATACAACAATGATAAATGGT
ATTGGCGTACTAGGATGGGGTGTTGGTGGTATTGAAGCTGAAGCTGGAATGCTTGGACAA
CCTTCTTATTTCCCAATTCCAGAGGTTATTGGTGTACGACTAGTAAATTCATTACCACAA
GGCGCAACAGCAACTGATTTAGCGTTAAGAGTAACTCAAGAGCTACGTAAAAAAGGTGTT
GTTGGTAAATTTGTGGAGTTCTTTGGTCCAGGTGTACAACATTTACCACTAGCAGACCGT
GCTACAATTGCAAACATGGCACCAGAGTATGGAGCAACTTGCGGATTCTTCCCAGTTGAT
GATGAATCTCTTAAATATATGAAGTTAACTGGTAGATCAGACGAACATATCGCGCTAGTA
AAAGAATATTTGAAACAAAACCATATGTTCTTTGATGTTGAGAAAGAAGATCCTAATTAT
ACAGATGTTATCGAATTGGATTTATCAACAGTTGAAGCATCGCTTTCAGGACCAAAACGT
CCTCAAGATTTAATTTTCTTAAGTGATATGAAATCATCATTTGAAAATTCTGTAACAGCT
CCAGCAGGCAACCAAGGACACGGTTTAGATAAAAGTGAATTTGATAAGAAAGCTGAAATT
AACTTTAAAGATGGATCAAAAGCTACAATGAAAACAGGTGATATTGCAATAGCAGCAATT
ACATCATGTACAAATACATCTAACCCTTATGTAATGTTAGGTGCAGGTTTAGTTGCTAAA
AAAGCAGTTGAAAAAGGCTTGAAAGTTCCTGAATACGTTAAAACTTCTCTAGCACCAGGA
TCAAAAGTTGTTACCGGATATTTAAGAGATGCTGGCTTACAACCTTATTTAGATGATTTA
GGCTTCAACTTGGTTGGTTATGGATGTACAACTTGTATCGGTAATTCAGGTCCTTTATTA
CCAGAAATTGAAAAAGCGATTGCTGATGAGGACCTATTAGTGACATCTGTATTATCTGGT
AACCGTAACTTTGAAGGTCGTATCCATCCTCTTGTTAAAGCCAATTACCTAGCTTCACCA
CAGTTAGTTGTTGCTTATGCATTAGCTGGAACGGTTGATATTGATTTACAAAATGAACCT
ATTGGTAAAGGTAATGACGGTGAAGATGTATATTTGAAAGATATTTGGCCATCAATTAAA
GAAGTTTCAGATACCGTTGATAGTGTTGTAACACCTGAATTATTTATTGAAGAATATAAT
AACGTATACAATAACAACGAATTATGGAATGAGATTGATGTAACTGATCAACCTCTATAT
GACTTTGATCCTAATTCAACATACATTCAAAATCCATCATTCTTCCAAGGATTATCTAAA
GAACCGGGTACGATTGTTCCATTAAATGGTTTACGTGTTATGGGTAAATTCGGTGATTCT
GTGACAACTGACCACATCTCTCCAGCAGGTGCAATTGGTAAAGATACGCCAGCTGGTAAA
TATTTACAAGATCATCAAGTGCCTATTCGTGAATTTAATTCATATGGTTCAAGACGTGGT
AATCACGAAGTAATGGTTCGAGGTACGTTTGCTAATATACGTATTAAAAACCAATTAGCG
CCAGGTACTGAAGGTGGTTTTACAACTTATTGGCCAACAAATGAAGTAATGCCTATCTTT
GATGCTGCAATGAAATATAAAGAAGATGGTACAGGTTTAGTTGTATTAGCTGGTAACGAT
TATGGTATGGGTTCATCTCGTGACTGGGCAGCAAAAGGTACAAACTTATTAGGTGTTAAA
ACAGTTATTGCACAAAGTTATGAACGTATCCATCGTTCAAATTTAGTTATGATGGGTGTA
TTACCATTAGAGTTTAAAAAAGGTGAATCAGCTGATTCTCTTGGTCTAGATGGTACAGAA
GAAATTTCTGTTAATATTGATGAAAATGTTCAACCACATGACTACGTCAAAGTTACTGCT
AAGAAGCAAGATGGTGATTTGGTAGAATTTGACGCTATGGTTCGTTTTGACTCACTTGTT
GAAATGGATTACTATCGTCACGGTGGAATTTTACAAATGGTTTTAAGAAATAAATTAGCG
CAATAA60
120
180
240
300
360
420
480
540
600
660
720
780
840
900
960
1020
1080
1140
1200
1260
1320
1380
1440
1500
1560
1620
1680
1740
1800
1860
1920
1980
2040
2100
2160
2220
2280
2340
2400
2460
2520
2580
2640
2700
2706
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SACOL1385 [new locus tag: SACOL_RS07050 ]
- symbol: AcnA
- description: aconitate hydratase
- length: 901
- theoretical pI: 4.58662
- theoretical MW: 98969
- GRAVY: -0.276249
⊟Function[edit | edit source]
- reaction: EC 4.2.1.3? ExPASyAconitate hydratase Citrate = isocitrateEC 4.2.1.99? ExPASy2-methylisocitrate dehydratase (2S,3R)-3-hydroxybutane-1,2,3-tricarboxylate = (Z)-but-2-ene-1,2,3-tricarboxylate + H2O
- TIGRFAM: Energy metabolism TCA cycle aconitate hydratase 1 (TIGR01341; EC 4.2.1.3; HMM-score: 1440.3)and 11 more2-methylisocitrate dehydratase, Fe/S-dependent (TIGR02333; EC 4.2.1.99; HMM-score: 785.6)Energy metabolism TCA cycle aconitate hydratase, mitochondrial (TIGR01340; EC 4.2.1.3; HMM-score: 240.6)Energy metabolism TCA cycle putative aconitate hydratase (TIGR01342; EC 4.2.1.3; HMM-score: 239.1)homoaconitate hydratase family protein (TIGR01343; HMM-score: 118.3)3-isopropylmalate dehydratase, large subunit (TIGR02086; HMM-score: 113.8)Amino acid biosynthesis Pyruvate family 3-isopropylmalate dehydratase, large subunit (TIGR02083; EC 4.2.1.33; HMM-score: 110.8)Amino acid biosynthesis Aspartate family homoaconitase (TIGR00139; EC 4.2.1.36; HMM-score: 97.8)Amino acid biosynthesis Pyruvate family 3-isopropylmalate dehydratase, large subunit (TIGR00170; EC 4.2.1.33; HMM-score: 89.8)3-isopropylmalate dehydratase, small subunit (TIGR02087; HMM-score: 48.5)Amino acid biosynthesis Pyruvate family 3-isopropylmalate dehydratase, small subunit (TIGR02084; EC 4.2.1.33; HMM-score: 37.5)Amino acid biosynthesis Pyruvate family 3-isopropylmalate dehydratase, small subunit (TIGR00171; EC 4.2.1.33; HMM-score: 27.8)
- TheSEED :
- Aconitate hydratase (EC 4.2.1.3)
and 2 more - PFAM: no clan defined Aconitase; Aconitase family (aconitate hydratase) (PF00330; HMM-score: 546.1)and 2 moreLeu-IlvD (CL0364) Aconitase_C; Aconitase C-terminal domain (PF00694; HMM-score: 141.2)no clan defined Bromodomain; Bromodomain (PF00439; HMM-score: 12.3)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors: [4Fe-4S] cluster
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 9.97
- Cytoplasmic Membrane Score: 0
- Cellwall Score: 0.01
- Extracellular Score: 0.02
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.7097
- Cytoplasmic Membrane Score: 0.0291
- Cell wall & surface Score: 0.0008
- Extracellular Score: 0.2604
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.001879
- TAT(Tat/SPI): 0.000137
- LIPO(Sec/SPII): 0.000282
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MAANFKEQSKKHFDLNGQSYTYYDLKAVEEQGITKVSNLPYSIRVLLESLLRQEDDFVITDDHIKALSQFGKDGNEGEVPFKPSRVILQDFTGVPAVVDLASLRKAMDDVGGDITKINPEVPVDLVIDHSVQVDSYANPEALERNMKLEFERNYERYQFLNWATKAFDNYNAVPPATGIVHQVNLEYLASVVHVRDVDGEKTAFPDTLVGTDSHTTMINGIGVLGWGVGGIEAEAGMLGQPSYFPIPEVIGVRLVNSLPQGATATDLALRVTQELRKKGVVGKFVEFFGPGVQHLPLADRATIANMAPEYGATCGFFPVDDESLKYMKLTGRSDEHIALVKEYLKQNHMFFDVEKEDPNYTDVIELDLSTVEASLSGPKRPQDLIFLSDMKSSFENSVTAPAGNQGHGLDKSEFDKKAEINFKDGSKATMKTGDIAIAAITSCTNTSNPYVMLGAGLVAKKAVEKGLKVPEYVKTSLAPGSKVVTGYLRDAGLQPYLDDLGFNLVGYGCTTCIGNSGPLLPEIEKAIADEDLLVTSVLSGNRNFEGRIHPLVKANYLASPQLVVAYALAGTVDIDLQNEPIGKGNDGEDVYLKDIWPSIKEVSDTVDSVVTPELFIEEYNNVYNNNELWNEIDVTDQPLYDFDPNSTYIQNPSFFQGLSKEPGTIVPLNGLRVMGKFGDSVTTDHISPAGAIGKDTPAGKYLQDHQVPIREFNSYGSRRGNHEVMVRGTFANIRIKNQLAPGTEGGFTTYWPTNEVMPIFDAAMKYKEDGTGLVVLAGNDYGMGSSRDWAAKGTNLLGVKTVIAQSYERIHRSNLVMMGVLPLEFKKGESADSLGLDGTEEISVNIDENVQPHDYVKVTAKKQDGDLVEFDAMVRFDSLVEMDYYRHGGILQMVLRNKLAQ
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas
- protein localization: Cytoplasmic [1] [2] [3] [4]
- quantitative data / protein copy number per cell: 5443 [5]
- interaction partners:
SACOL1760 (ackA) acetate kinase [6] (data from MRSA252) SACOL0660 (adhP) alcohol dehydrogenase [6] (data from MRSA252) SACOL2218 (adk) adenylate kinase [6] (data from MRSA252) SACOL0452 (ahpC) alkyl hydroperoxide reductase subunit C [6] (data from MRSA252) SACOL1758 (ald2) alanine dehydrogenase [6] (data from MRSA252) SACOL2656 (arcB2) ornithine carbamoyltransferase [6] (data from MRSA252) SACOL0663 (argS) arginyl-tRNA synthetase [6] (data from MRSA252) SACOL1494 (asnC) asparaginyl-tRNA synthetase [6] (data from MRSA252) SACOL1215 (carB) carbamoyl phosphate synthase large subunit [6] (data from MRSA252) SACOL1786 (ccpA) catabolite control protein A [6] (data from MRSA252) SACOL0833 (clpP) ATP-dependent Clp protease proteolytic subunit [6] (data from MRSA252) SACOL0557 (cysK) cysteine synthase [6] (data from MRSA252) SACOL1800 (dat) D-alanine aminotransferase [6] (data from MRSA252) SACOL2130 (deoD) purine nucleoside phosphorylase [6] (data from MRSA252) SACOL1637 (dnaK) molecular chaperone DnaK [6] (data from MRSA252) SACOL0002 (dnaN) DNA polymerase III subunit beta [6] (data from MRSA252) SACOL0634 (eutD) phosphotransacetylase [6] (data from MRSA252) SACOL0988 (fabF) 3-oxoacyl-ACP synthase [6] (data from MRSA252) SACOL1016 (fabI) enoyl-ACP reductase [6] (data from MRSA252) SACOL2091 (fabZ) (3R)-hydroxymyristoyl-ACP dehydratase [6] (data from MRSA252) SACOL2117 (fbaA) fructose-bisphosphate aldolase [6] (data from MRSA252) SACOL2622 (fdaB) fructose-1,6-bisphosphate aldolase [6] (data from MRSA252) SACOL1329 (femC) glutamine synthetase [6] (data from MRSA252) SACOL1782 (fhs) formate--tetrahydrofolate ligase [6] (data from MRSA252) SACOL1072 (folD) bifunctional 5,10-methylene-tetrahydrofolate dehydrogenase/ 5,10-methylene-tetrahydrofolate cyclohydrolase [6] (data from MRSA252) SACOL1198 (ftsA) cell division protein FtsA [6] (data from MRSA252) SACOL0593 (fusA) elongation factor G [6] (data from MRSA252) SACOL0838 (gapA1) glyceraldehyde 3-phosphate dehydrogenase [6] (data from MRSA252) SACOL1960 (gatB) aspartyl/glutamyl-tRNA amidotransferase subunit B [6] (data from MRSA252) SACOL0877 (gcvH) glycine cleavage system protein H [6] (data from MRSA252) SACOL2151 (glmM) phosphoglucosamine mutase [6] (data from MRSA252) SACOL2145 (glmS) glucosamine--fructose-6-phosphate aminotransferase [6] (data from MRSA252) SACOL1742 (gltA) citrate synthase [6] (data from MRSA252) SACOL0961 (gluD) glutamate dehydrogenase [6] (data from MRSA252) SACOL1622 (glyS) glycyl-tRNA synthetase [6] (data from MRSA252) SACOL1554 (gnd) 6-phosphogluconate dehydrogenase [6] (data from MRSA252) SACOL1665 (greA) transcription elongation factor GreA [6] (data from MRSA252) SACOL2016 (groEL) chaperonin GroEL [6] (data from MRSA252) SACOL2017 (groES) co-chaperonin GroES [6] (data from MRSA252) SACOL0460 (guaB) inosine-5'-monophosphate dehydrogenase [6] (data from MRSA252) SACOL0554 (hpt) hypoxanthine phosphoribosyltransferase [6] (data from MRSA252) SACOL1513 (hup) DNA-binding protein HU [6] (data from MRSA252) SACOL1206 (ileS) isoleucyl-tRNA synthetase [6] (data from MRSA252) SACOL0600 (ilvE) branched-chain amino acid aminotransferase [6] (data from MRSA252) SACOL0240 (ispD) 2-C-methyl-D-erythritol 4-phosphate cytidylyltransferase [6] (data from MRSA252) SACOL1368 (kataA) catalase [6] (data from MRSA252) SACOL0222 (ldh1) L-lactate dehydrogenase [6] (data from MRSA252) SACOL2618 (ldh2) L-lactate dehydrogenase [6] (data from MRSA252) SACOL0562 (lysS) lysyl-tRNA synthetase [6] (data from MRSA252) SACOL1054 (menB) naphthoate synthase [6] (data from MRSA252) SACOL2623 (mqo2) malate:quinone oxidoreductase [6] (data from MRSA252) SACOL2116 (murAB) UDP-N-acetylglucosamine 1-carboxyvinyltransferase [6] (data from MRSA252) SACOL1509 (ndk) nucleoside diphosphate kinase [6] (data from MRSA252) SACOL0746 (norR) MarR family transcriptional regulator [6] (data from MRSA252) SACOL0792 (nrdE) ribonucleotide-diphosphate reductase subunit alpha [6] (data from MRSA252) SACOL0793 (nrdF) ribonucleotide-diphosphate reductase subunit beta [6] (data from MRSA252) SACOL1285 (nusA) transcription elongation factor NusA [6] (data from MRSA252) SACOL2128 (pdp) pyrimidine-nucleoside phosphorylase [6] (data from MRSA252) SACOL1005 (pepF) oligoendopeptidase F [6] (data from MRSA252) SACOL1746 (pfkA) 6-phosphofructokinase [6] (data from MRSA252) SACOL0204 (pflB) formate acetyltransferase [6] (data from MRSA252) SACOL0966 (pgi) glucose-6-phosphate isomerase [6] (data from MRSA252) SACOL0839 (pgk) phosphoglycerate kinase [6] (data from MRSA252) SACOL0841 (pgm) phosphoglyceromutase [6] (data from MRSA252) SACOL1293 (pnp) polynucleotide phosphorylase/polyadenylase [6] (data from MRSA252) SACOL1982 (ppaC) manganese-dependent inorganic pyrophosphatase [6] (data from MRSA252) SACOL1091 (ptsH) phosphocarrier protein HPr [6] (data from MRSA252) SACOL1092 (ptsI) phosphoenolpyruvate-protein phosphotransferase [6] (data from MRSA252) SACOL1745 (pyk) pyruvate kinase [6] (data from MRSA252) SACOL0584 (rplA) 50S ribosomal protein L1 [6] (data from MRSA252) SACOL2227 (rplE) 50S ribosomal protein L5 [6] (data from MRSA252) SACOL2224 (rplF) 50S ribosomal protein L6 [6] (data from MRSA252) SACOL0585 (rplJ) 50S ribosomal protein L10 [6] (data from MRSA252) SACOL0583 (rplK) 50S ribosomal protein L11 [6] (data from MRSA252) SACOL2220 (rplO) 50S ribosomal protein L15 [6] (data from MRSA252) SACOL2212 (rplQ) 50S ribosomal protein L17 [6] (data from MRSA252) SACOL1725 (rplT) 50S ribosomal protein L20 [6] (data from MRSA252) SACOL1702 (rplU) 50S ribosomal protein L21 [6] (data from MRSA252) SACOL2237 (rplW) 50S ribosomal protein L23 [6] (data from MRSA252) SACOL0545 (rplY) 50S ribosomal protein L25/general stress protein Ctc [6] (data from MRSA252) SACOL2221 (rpmD) 50S ribosomal protein L30 [6] (data from MRSA252) SACOL2213 (rpoA) DNA-directed RNA polymerase subunit alpha [6] (data from MRSA252) SACOL0589 (rpoC) DNA-directed RNA polymerase subunit beta' [6] (data from MRSA252) SACOL2120 (rpoE) DNA-directed RNA polymerase subunit delta [6] (data from MRSA252) SACOL1516 (rpsA) 30S ribosomal protein S1 [6] (data from MRSA252) SACOL1274 (rpsB) 30S ribosomal protein S2 [6] (data from MRSA252) SACOL2233 (rpsC) 30S ribosomal protein S3 [6] (data from MRSA252) SACOL1769 (rpsD) 30S ribosomal protein S4 [6] (data from MRSA252) SACOL0592 (rpsG) 30S ribosomal protein S7 [6] (data from MRSA252) SACOL2235 (rpsS) 30S ribosomal protein S19 [6] (data from MRSA252) SACOL2056 (rsbV) anti-anti-sigma factor RsbV [6] (data from MRSA252) SACOL2055 (rsbW) serine-protein kinase RsbW [6] (data from MRSA252) SACOL0009 (serS) seryl-tRNA synthetase [6] (data from MRSA252) SACOL1610 (sodA2) superoxide dismutase [6] (data from MRSA252) SACOL0095 (spa) immunoglobulin G binding protein A precursor [6] (data from MRSA252) SACOL1262 (sucC) succinyl-CoA synthetase subunit beta [6] (data from MRSA252) SACOL1831 (tal) translaldolase [6] (data from MRSA252) SACOL1729 (thrS) threonyl-tRNA synthetase [6] (data from MRSA252) SACOL1722 (tig) trigger factor [6] (data from MRSA252) SACOL1377 (tkt) transketolase [6] (data from MRSA252) SACOL1762 (tpx) thiol peroxidase [6] (data from MRSA252) SACOL1155 (trxA) thioredoxin [6] (data from MRSA252) SACOL1276 (tsf) elongation factor Ts [6] (data from MRSA252) SACOL0594 (tuf) elongation factor Tu [6] (data from MRSA252) SACOL2104 (upp) uracil phosphoribosyltransferase [6] (data from MRSA252) SACOL0457 hypothetical protein [6] (data from MRSA252) SACOL0564 pyridoxal biosynthesis lyase PdxS [6] (data from MRSA252) SACOL0596 2-amino-3-ketobutyrate coenzyme A ligase [6] (data from MRSA252) SACOL0688 ABC transporter substrate-binding protein [6] (data from MRSA252) SACOL0815 ribosomal subunit interface protein [6] (data from MRSA252) SACOL0876 hypothetical protein [6] (data from MRSA252) SACOL0944 NADH dehydrogenase [6] (data from MRSA252) SACOL1020 hypothetical protein [6] (data from MRSA252) SACOL1427 ABC transporter ATP-binding protein [6] (data from MRSA252) SACOL1454 [6] (data from MRSA252) SACOL1457 PTS system, IIA component [6] (data from MRSA252) SACOL1593 glycine dehydrogenase subunit 2 [6] (data from MRSA252) SACOL1801 dipeptidase PepV [6] (data from MRSA252) SACOL1952 ferritins family protein [6] (data from MRSA252) SACOL2131 Dps family protein [6] (data from MRSA252) SACOL2173 alkaline shock protein 23 [6] (data from MRSA252) SACOL2296 glycerate dehydrogenase [6] (data from MRSA252) SACOL2367 alcohol dehydrogenase, zinc-containing [6] (data from MRSA252) SACOL2535 D-lactate dehydrogenase [6] (data from MRSA252) SACOL2561 hydroxymethylglutaryl-CoA synthase [6] (data from MRSA252) SACOL2569 1-pyrroline-5-carboxylate dehydrogenase [6] (data from MRSA252)
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: no polycistronic organisation predicted
⊟Regulation[edit | edit source]
- regulators: Fur* (repression) regulon, CcpA regulon
Fur* (TF) important in Iron homeostasis; RegPrecise CcpA (TF) important in Carbon catabolism; RegPrecise
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: data available for NCTC8325
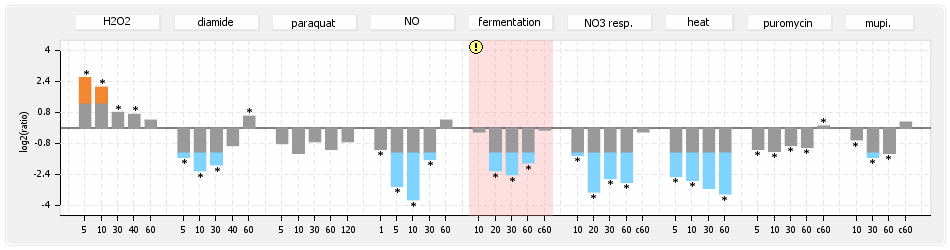
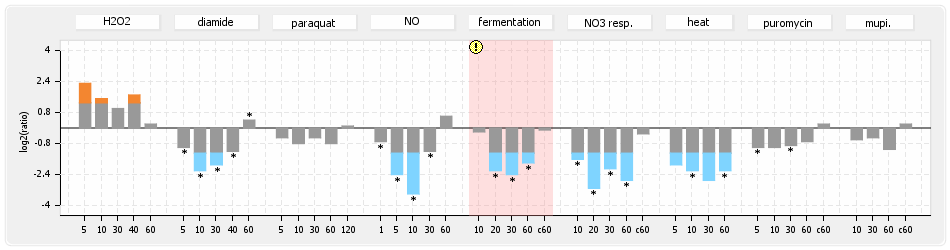
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Dörte Becher, Kristina Hempel, Susanne Sievers, Daniela Zühlke, Jan Pané-Farré, Andreas Otto, Stephan Fuchs, Dirk Albrecht, Jörg Bernhardt, Susanne Engelmann, Uwe Völker, Jan Maarten van Dijl, Michael Hecker
A proteomic view of an important human pathogen--towards the quantification of the entire Staphylococcus aureus proteome.
PLoS One: 2009, 4(12);e8176
[PubMed:19997597] [WorldCat.org] [DOI] (I e) - ↑ Kristina Hempel, Jan Pané-Farré, Andreas Otto, Susanne Sievers, Michael Hecker, Dörte Becher
Quantitative cell surface proteome profiling for SigB-dependent protein expression in the human pathogen Staphylococcus aureus via biotinylation approach.
J Proteome Res: 2010, 9(3);1579-90
[PubMed:20108986] [WorldCat.org] [DOI] (I p) - ↑ Kristina Hempel, Florian-Alexander Herbst, Martin Moche, Michael Hecker, Dörte Becher
Quantitative proteomic view on secreted, cell surface-associated, and cytoplasmic proteins of the methicillin-resistant human pathogen Staphylococcus aureus under iron-limited conditions.
J Proteome Res: 2011, 10(4);1657-66
[PubMed:21323324] [WorldCat.org] [DOI] (I p) - ↑ Andreas Otto, Jan Maarten van Dijl, Michael Hecker, Dörte Becher
The Staphylococcus aureus proteome.
Int J Med Microbiol: 2014, 304(2);110-20
[PubMed:24439828] [WorldCat.org] [DOI] (I p) - ↑ Daniela Zühlke, Kirsten Dörries, Jörg Bernhardt, Sandra Maaß, Jan Muntel, Volkmar Liebscher, Jan Pané-Farré, Katharina Riedel, Michael Lalk, Uwe Völker, Susanne Engelmann, Dörte Becher, Stephan Fuchs, Michael Hecker
Costs of life - Dynamics of the protein inventory of Staphylococcus aureus during anaerobiosis.
Sci Rep: 2016, 6;28172
[PubMed:27344979] [WorldCat.org] [DOI] (I e) - ↑ 6.000 6.001 6.002 6.003 6.004 6.005 6.006 6.007 6.008 6.009 6.010 6.011 6.012 6.013 6.014 6.015 6.016 6.017 6.018 6.019 6.020 6.021 6.022 6.023 6.024 6.025 6.026 6.027 6.028 6.029 6.030 6.031 6.032 6.033 6.034 6.035 6.036 6.037 6.038 6.039 6.040 6.041 6.042 6.043 6.044 6.045 6.046 6.047 6.048 6.049 6.050 6.051 6.052 6.053 6.054 6.055 6.056 6.057 6.058 6.059 6.060 6.061 6.062 6.063 6.064 6.065 6.066 6.067 6.068 6.069 6.070 6.071 6.072 6.073 6.074 6.075 6.076 6.077 6.078 6.079 6.080 6.081 6.082 6.083 6.084 6.085 6.086 6.087 6.088 6.089 6.090 6.091 6.092 6.093 6.094 6.095 6.096 6.097 6.098 6.099 6.100 6.101 6.102 6.103 6.104 6.105 6.106 6.107 6.108 6.109 6.110 6.111 6.112 6.113 6.114 6.115 6.116 6.117 6.118 6.119 6.120 6.121 6.122 6.123 6.124 6.125 Artem Cherkasov, Michael Hsing, Roya Zoraghi, Leonard J Foster, Raymond H See, Nikolay Stoynov, Jihong Jiang, Sukhbir Kaur, Tian Lian, Linda Jackson, Huansheng Gong, Rick Swayze, Emily Amandoron, Farhad Hormozdiari, Phuong Dao, Cenk Sahinalp, Osvaldo Santos-Filho, Peter Axerio-Cilies, Kendall Byler, William R McMaster, Robert C Brunham, B Brett Finlay, Neil E Reiner
Mapping the protein interaction network in methicillin-resistant Staphylococcus aureus.
J Proteome Res: 2011, 10(3);1139-50
[PubMed:21166474] [WorldCat.org] [DOI] (I p)