Jump to navigation
Jump to search
NCBI: 10-JUN-2013
⊟Summary[edit | edit source]
- organism: Staphylococcus aureus COL
- locus tag: SACOL1277 [new locus tag: SACOL_RS06520 ]
- pan locus tag?: SAUPAN003558000
- symbol: pyrH
- pan gene symbol?: pyrH
- synonym:
- product: uridylate kinase
⊟Genome View[edit | edit source]
⊟Gene[edit | edit source]
⊟General[edit | edit source]
- type: CDS
- locus tag: SACOL1277 [new locus tag: SACOL_RS06520 ]
- symbol: pyrH
- product: uridylate kinase
- replicon: chromosome
- strand: +
- coordinates: 1286562..1287284
- length: 723
- essential: unknown other strains
⊟Accession numbers[edit | edit source]
- Gene ID: 3238508 NCBI
- RefSeq: YP_186134 NCBI
- BioCyc: see SACOL_RS06520
- MicrobesOnline: 912740 MicrobesOnline
⊟Phenotype[edit | edit source]
Share your knowledge and add information here. [edit]
⊟DNA sequence[edit | edit source]
- 1
61
121
181
241
301
361
421
481
541
601
661
721ATGGCTCAAATTTCTAAATATAAACGTGTAGTTTTGAAACTAAGTGGTGAAGCGTTAGCT
GGAGAAAAAGGATTTGGCATAAATCCAGTAATTATTAAAAGTGTTGCTGAGCAAGTGGCT
GAAGTTGCTAAAATGGACTGTGAAATCGCAGTAATCGTTGGTGGCGGAAACATTTGGAGA
GGTAAAACAGGTAGTGACTTAGGTATGGACCGTGGAACTGCTGATTACATGGGTATGCTT
GCAACTGTAATGAATGCCTTAGCATTACAAGATAGTTTAGAACAATTGGATTGTGATACA
CGAGTATTAACATCTATTGAAATGAAGCAAGTGGCTGAACCTTATATTCGTCGTCGTGCA
ATTAGACACTTAGAAAAGAAACGCGTAGTTATTTTTGCTGCAGGTATTGGAAACCCATAC
TTCTCTACAGATACTACAGCGGCATTACGTGCTGCAGAAGTTGAAGCAGATGTTATTTTA
ATGGGCAAAAATAATGTAGATGGTGTATATTCTGCAGATCCTAAAGTAAACAAAGATGCG
GTAAAATATGAACATTTAACGCATATTCAAATGCTTCAAGAAGGTTTACAAGTAATGGAT
TCAACAGCATCCTCATTCTGTATGGATAATAACATTCCGTTAACTGTTTTCTCTATTATG
GAAGAAGGAAATATTAAACGTGCTGTTATGGGTGAAAAGATAGGTACGTTAATTACAAAA
TAA60
120
180
240
300
360
420
480
540
600
660
720
723
⊟Protein[edit | edit source]
⊟General[edit | edit source]
- locus tag: SACOL1277 [new locus tag: SACOL_RS06520 ]
- symbol: PyrH
- description: uridylate kinase
- length: 240
- theoretical pI: 6.28636
- theoretical MW: 26145.2
- GRAVY: -0.00458333
⊟Function[edit | edit source]
- reaction: EC 2.7.4.22? ExPASyUMP kinase ATP + UMP = ADP + UDP
- TIGRFAM: Purines, pyrimidines, nucleosides, and nucleotides Nucleotide and nucleoside interconversions UMP kinase (TIGR02075; EC 2.7.4.22; HMM-score: 345.7)and 4 morePurines, pyrimidines, nucleosides, and nucleotides Nucleotide and nucleoside interconversions putative uridylate kinase (TIGR02076; EC 2.7.4.-; HMM-score: 93)Amino acid biosynthesis Aspartate family aspartate kinase (TIGR00657; EC 2.7.2.4; HMM-score: 37)Amino acid biosynthesis Aspartate family aspartate kinase, monofunctional class (TIGR00656; EC 2.7.2.4; HMM-score: 32)Amino acid biosynthesis Glutamate family glutamate 5-kinase (TIGR01027; EC 2.7.2.11; HMM-score: 27.5)
- TheSEED :
- Uridylate kinase (EC 2.7.4.22)
- PFAM: no clan defined AA_kinase; Amino acid kinase family (PF00696; HMM-score: 117.8)
⊟Structure, modifications & cofactors[edit | edit source]
- domains:
- modifications:
- cofactors:
- effectors:
⊟Localization[edit | edit source]
- PSORTb: Cytoplasmic
- Cytoplasmic Score: 7.5
- Cytoplasmic Membrane Score: 1.15
- Cellwall Score: 0.62
- Extracellular Score: 0.73
- Internal Helices: 0
- DeepLocPro: Cytoplasmic
- Cytoplasmic Score: 0.9446
- Cytoplasmic Membrane Score: 0.0189
- Cell wall & surface Score: 0
- Extracellular Score: 0.0365
- LocateP: Intracellular
- Prediction by SwissProt Classification: Cytoplasmic
- Pathway Prediction: No pathway
- Intracellular possibility: 1
- Signal peptide possibility: -1
- N-terminally Anchored Score: 1
- Predicted Cleavage Site: No CleavageSite
- SignalP: no predicted signal peptide
- SP(Sec/SPI): 0.013405
- TAT(Tat/SPI): 0.000561
- LIPO(Sec/SPII): 0.001311
- predicted transmembrane helices (TMHMM): 0
⊟Accession numbers[edit | edit source]
⊟Protein sequence[edit | edit source]
- MAQISKYKRVVLKLSGEALAGEKGFGINPVIIKSVAEQVAEVAKMDCEIAVIVGGGNIWRGKTGSDLGMDRGTADYMGMLATVMNALALQDSLEQLDCDTRVLTSIEMKQVAEPYIRRRAIRHLEKKRVVIFAAGIGNPYFSTDTTAALRAAEVEADVILMGKNNVDGVYSADPKVNKDAVKYEHLTHIQMLQEGLQVMDSTASSFCMDNNIPLTVFSIMEEGNIKRAVMGEKIGTLITK
⊟Experimental data[edit | edit source]
- experimentally validated: PeptideAtlas
- protein localization: Cytoplasmic [1] [2] [3] [4]
- quantitative data / protein copy number per cell: 344 [5]
- interaction partners:
SACOL1760 (ackA) acetate kinase [6] (data from MRSA252) SACOL1385 (acnA) aconitate hydratase [6] (data from MRSA252) SACOL0660 (adhP) alcohol dehydrogenase [6] (data from MRSA252) SACOL2218 (adk) adenylate kinase [6] (data from MRSA252) SACOL0452 (ahpC) alkyl hydroperoxide reductase subunit C [6] (data from MRSA252) SACOL0451 (ahpF) alkyl hydroperoxide reductase subunit F [6] (data from MRSA252) SACOL1673 (alaS) alanyl-tRNA synthetase [6] (data from MRSA252) SACOL1478 (ald1) alanine dehydrogenase [6] (data from MRSA252) SACOL1758 (ald2) alanine dehydrogenase [6] (data from MRSA252) SACOL0154 (aldA1) aldehyde dehydrogenase [6] (data from MRSA252) SACOL2657 (arcA) arginine deiminase [6] (data from MRSA252) SACOL2656 (arcB2) ornithine carbamoyltransferase [6] (data from MRSA252) SACOL2654 (arcC2) carbamate kinase [6] (data from MRSA252) SACOL0663 (argS) arginyl-tRNA synthetase [6] (data from MRSA252) SACOL1494 (asnC) asparaginyl-tRNA synthetase [6] (data from MRSA252) SACOL1685 (aspS) aspartyl-tRNA synthetase [6] (data from MRSA252) SACOL1215 (carB) carbamoyl phosphate synthase large subunit [6] (data from MRSA252) SACOL1786 (ccpA) catabolite control protein A [6] (data from MRSA252) SACOL0570 (clpC) [6] (data from MRSA252) SACOL0833 (clpP) ATP-dependent Clp protease proteolytic subunit [6] (data from MRSA252) SACOL1721 (clpX) ATP-dependent protease ATP-binding subunit ClpX [6] (data from MRSA252) SACOL1518 (cmk) cytidylate kinase [6] (data from MRSA252) SACOL1272 (codY) transcriptional repressor CodY [6] (data from MRSA252) SACOL0557 (cysK) cysteine synthase [6] (data from MRSA252) SACOL1800 (dat) D-alanine aminotransferase [6] (data from MRSA252) SACOL2074 (ddl) D-alanyl-alanine synthetase A [6] (data from MRSA252) SACOL1100 (def) peptide deformylase [6] (data from MRSA252) SACOL0124 (deoB) phosphopentomutase [6] (data from MRSA252) SACOL0123 (deoC1) deoxyribose-phosphate aldolase [6] (data from MRSA252) SACOL2130 (deoD) purine nucleoside phosphorylase [6] (data from MRSA252) SACOL0935 (dltA) D-alanine--poly(phosphoribitol) ligase subunit 1 [6] (data from MRSA252) SACOL0937 (dltC) D-alanine--poly(phosphoribitol) ligase subunit 2 [6] (data from MRSA252) SACOL1637 (dnaK) molecular chaperone DnaK [6] (data from MRSA252) SACOL0002 (dnaN) DNA polymerase III subunit beta [6] (data from MRSA252) SACOL1587 (efp) elongation factor P [6] (data from MRSA252) SACOL0842 (eno) phosphopyruvate hydratase [6] (data from MRSA252) SACOL0634 (eutD) phosphotransacetylase [6] (data from MRSA252) SACOL1244 (fabD) malonyl CoA-ACP transacylase [6] (data from MRSA252) SACOL0988 (fabF) 3-oxoacyl-ACP synthase [6] (data from MRSA252) SACOL1245 (fabG1) 3-oxoacyl-ACP reductase [6] (data from MRSA252) SACOL0987 (fabH) 3-oxoacyl-ACP synthase [6] (data from MRSA252) SACOL1016 (fabI) enoyl-ACP reductase [6] (data from MRSA252) SACOL2091 (fabZ) (3R)-hydroxymyristoyl-ACP dehydratase [6] (data from MRSA252) SACOL2117 (fbaA) fructose-bisphosphate aldolase [6] (data from MRSA252) SACOL2622 (fdaB) fructose-1,6-bisphosphate aldolase [6] (data from MRSA252) SACOL1329 (femC) glutamine synthetase [6] (data from MRSA252) SACOL1782 (fhs) formate--tetrahydrofolate ligase [6] (data from MRSA252) SACOL1072 (folD) bifunctional 5,10-methylene-tetrahydrofolate dehydrogenase/ 5,10-methylene-tetrahydrofolate cyclohydrolase [6] (data from MRSA252) SACOL1278 (frr) ribosome recycling factor [6] (data from MRSA252) SACOL1198 (ftsA) cell division protein FtsA [6] (data from MRSA252) SACOL1199 (ftsZ) cell division protein FtsZ [6] (data from MRSA252) SACOL0593 (fusA) elongation factor G [6] (data from MRSA252) SACOL0838 (gapA1) glyceraldehyde 3-phosphate dehydrogenase [6] (data from MRSA252) SACOL1734 (gapA2) glyceraldehyde 3-phosphate dehydrogenase 2 [6] (data from MRSA252) SACOL1961 (gatA) aspartyl/glutamyl-tRNA amidotransferase subunit A [6] (data from MRSA252) SACOL1960 (gatB) aspartyl/glutamyl-tRNA amidotransferase subunit B [6] (data from MRSA252) SACOL0877 (gcvH) glycine cleavage system protein H [6] (data from MRSA252) SACOL1595 (gcvT) glycine cleavage system aminomethyltransferase T [6] (data from MRSA252) SACOL2151 (glmM) phosphoglucosamine mutase [6] (data from MRSA252) SACOL2145 (glmS) glucosamine--fructose-6-phosphate aminotransferase [6] (data from MRSA252) SACOL1742 (gltA) citrate synthase [6] (data from MRSA252) SACOL0574 (gltX) glutamyl-tRNA synthetase [6] (data from MRSA252) SACOL0961 (gluD) glutamate dehydrogenase [6] (data from MRSA252) SACOL1622 (glyS) glycyl-tRNA synthetase [6] (data from MRSA252) SACOL1554 (gnd) 6-phosphogluconate dehydrogenase [6] (data from MRSA252) SACOL2415 (gpmA) phosphoglyceromutase [6] (data from MRSA252) SACOL1665 (greA) transcription elongation factor GreA [6] (data from MRSA252) SACOL2016 (groEL) chaperonin GroEL [6] (data from MRSA252) SACOL1638 (grpE) heat shock protein GrpE [6] (data from MRSA252) SACOL0461 (guaA) GMP synthase [6] (data from MRSA252) SACOL0460 (guaB) inosine-5'-monophosphate dehydrogenase [6] (data from MRSA252) SACOL0554 (hpt) hypoxanthine phosphoribosyltransferase [6] (data from MRSA252) SACOL1513 (hup) DNA-binding protein HU [6] (data from MRSA252) SACOL1741 (icd) isocitrate dehydrogenase [6] (data from MRSA252) SACOL1206 (ileS) isoleucyl-tRNA synthetase [6] (data from MRSA252) SACOL0600 (ilvE) branched-chain amino acid aminotransferase [6] (data from MRSA252) SACOL1288 (infB) translation initiation factor IF-2 [6] (data from MRSA252) SACOL1727 (infC) translation initiation factor IF-3 [6] (data from MRSA252) SACOL0240 (ispD) 2-C-methyl-D-erythritol 4-phosphate cytidylyltransferase [6] (data from MRSA252) SACOL1368 (kataA) catalase [6] (data from MRSA252) SACOL0222 (ldh1) L-lactate dehydrogenase [6] (data from MRSA252) SACOL2618 (ldh2) L-lactate dehydrogenase [6] (data from MRSA252) SACOL1808 (leuS) leucyl-tRNA synthetase [6] (data from MRSA252) SACOL0562 (lysS) lysyl-tRNA synthetase [6] (data from MRSA252) SACOL2664 (manA2) mannose-6-phosphate isomerase [6] (data from MRSA252) SACOL1054 (menB) naphthoate synthase [6] (data from MRSA252) SACOL1837 (metK) S-adenosylmethionine synthetase [6] (data from MRSA252) SACOL0533 (metS) methionyl-tRNA synthetase [6] (data from MRSA252) SACOL2623 (mqo2) malate:quinone oxidoreductase [6] (data from MRSA252) SACOL2149 (mtlD) mannitol-1-phosphate 5-dehydrogenase [6] (data from MRSA252) SACOL2092 (murAA) UDP-N-acetylglucosamine 1-carboxyvinyltransferase [6] (data from MRSA252) SACOL2116 (murAB) UDP-N-acetylglucosamine 1-carboxyvinyltransferase [6] (data from MRSA252) SACOL1790 (murC) UDP-N-acetylmuramate--L-alanine ligase [6] (data from MRSA252) SACOL1509 (ndk) nucleoside diphosphate kinase [6] (data from MRSA252) SACOL0746 (norR) MarR family transcriptional regulator [6] (data from MRSA252) SACOL0792 (nrdE) ribonucleotide-diphosphate reductase subunit alpha [6] (data from MRSA252) SACOL0793 (nrdF) ribonucleotide-diphosphate reductase subunit beta [6] (data from MRSA252) SACOL1285 (nusA) transcription elongation factor NusA [6] (data from MRSA252) SACOL0582 (nusG) transcription antitermination protein [6] (data from MRSA252) SACOL2615 (panB) 3-methyl-2-oxobutanoate hydroxymethyltransferase [6] (data from MRSA252) SACOL1838 (pckA) phosphoenolpyruvate carboxykinase [6] (data from MRSA252) SACOL1102 (pdhA) pyruvate dehydrogenase complex E1 component subunit alpha [6] (data from MRSA252) SACOL1103 (pdhB) pyruvate dehydrogenase complex E1 component subunit beta [6] (data from MRSA252) SACOL1104 (pdhC) branched-chain alpha-keto acid dehydrogenase E2 [6] (data from MRSA252) SACOL1105 (pdhD) dihydrolipoamide dehydrogenase [6] (data from MRSA252) SACOL2128 (pdp) pyrimidine-nucleoside phosphorylase [6] (data from MRSA252) SACOL1005 (pepF) oligoendopeptidase F [6] (data from MRSA252) SACOL1746 (pfkA) 6-phosphofructokinase [6] (data from MRSA252) SACOL0204 (pflB) formate acetyltransferase [6] (data from MRSA252) SACOL0966 (pgi) glucose-6-phosphate isomerase [6] (data from MRSA252) SACOL0839 (pgk) phosphoglycerate kinase [6] (data from MRSA252) SACOL0841 (pgm) phosphoglyceromutase [6] (data from MRSA252) SACOL1293 (pnp) polynucleotide phosphorylase/polyadenylase [6] (data from MRSA252) SACOL1982 (ppaC) manganese-dependent inorganic pyrophosphatase [6] (data from MRSA252) SACOL1282 (proS) prolyl-tRNA synthetase [6] (data from MRSA252) SACOL0544 (prsA) ribose-phosphate pyrophosphokinase [6] (data from MRSA252) SACOL1091 (ptsH) phosphocarrier protein HPr [6] (data from MRSA252) SACOL1092 (ptsI) phosphoenolpyruvate-protein phosphotransferase [6] (data from MRSA252) SACOL1816 (putA) proline dehydrogenase [6] (data from MRSA252) SACOL1745 (pyk) pyruvate kinase [6] (data from MRSA252) SACOL1213 (pyrC) dihydroorotase [6] (data from MRSA252) SACOL1217 (pyrE) orotate phosphoribosyltransferase [6] (data from MRSA252) SACOL2119 (pyrG) CTP synthetase [6] (data from MRSA252) SACOL0584 (rplA) 50S ribosomal protein L1 [6] (data from MRSA252) SACOL2236 (rplB) 50S ribosomal protein L2 [6] (data from MRSA252) SACOL2239 (rplC) 50S ribosomal protein L3 [6] (data from MRSA252) SACOL2238 (rplD) 50S ribosomal protein L4 [6] (data from MRSA252) SACOL2227 (rplE) 50S ribosomal protein L5 [6] (data from MRSA252) SACOL2224 (rplF) 50S ribosomal protein L6 [6] (data from MRSA252) SACOL0015 (rplI) 50S ribosomal protein L9 [6] (data from MRSA252) SACOL0585 (rplJ) 50S ribosomal protein L10 [6] (data from MRSA252) SACOL0583 (rplK) 50S ribosomal protein L11 [6] (data from MRSA252) SACOL0586 (rplL) 50S ribosomal protein L7/L12 [6] (data from MRSA252) SACOL2207 (rplM) 50S ribosomal protein L13 [6] (data from MRSA252) SACOL2220 (rplO) 50S ribosomal protein L15 [6] (data from MRSA252) SACOL2212 (rplQ) 50S ribosomal protein L17 [6] (data from MRSA252) SACOL1257 (rplS) 50S ribosomal protein L19 [6] (data from MRSA252) SACOL1725 (rplT) 50S ribosomal protein L20 [6] (data from MRSA252) SACOL1702 (rplU) 50S ribosomal protein L21 [6] (data from MRSA252) SACOL2234 (rplV) 50S ribosomal protein L22 [6] (data from MRSA252) SACOL2237 (rplW) 50S ribosomal protein L23 [6] (data from MRSA252) SACOL2228 (rplX) 50S ribosomal protein L24 [6] (data from MRSA252) SACOL0545 (rplY) 50S ribosomal protein L25/general stress protein Ctc [6] (data from MRSA252) SACOL2221 (rpmD) 50S ribosomal protein L30 [6] (data from MRSA252) SACOL2112 (rpmE2) 50S ribosomal protein L31 [6] (data from MRSA252) SACOL1726 (rpmI) 50S ribosomal protein L35 [6] (data from MRSA252) SACOL2213 (rpoA) DNA-directed RNA polymerase subunit alpha [6] (data from MRSA252) SACOL0588 (rpoB) DNA-directed RNA polymerase subunit beta [6] (data from MRSA252) SACOL0589 (rpoC) DNA-directed RNA polymerase subunit beta' [6] (data from MRSA252) SACOL1516 (rpsA) 30S ribosomal protein S1 [6] (data from MRSA252) SACOL1274 (rpsB) 30S ribosomal protein S2 [6] (data from MRSA252) SACOL2233 (rpsC) 30S ribosomal protein S3 [6] (data from MRSA252) SACOL1769 (rpsD) 30S ribosomal protein S4 [6] (data from MRSA252) SACOL2222 (rpsE) 30S ribosomal protein S5 [6] (data from MRSA252) SACOL0437 (rpsF) 30S ribosomal protein S6 [6] (data from MRSA252) SACOL0592 (rpsG) 30S ribosomal protein S7 [6] (data from MRSA252) SACOL2225 (rpsH) 30S ribosomal protein S8 [6] (data from MRSA252) SACOL2206 (rpsI) 30S ribosomal protein S9 [6] (data from MRSA252) SACOL2240 (rpsJ) 30S ribosomal protein S10 [6] (data from MRSA252) SACOL2214 (rpsK) 30S ribosomal protein S11 [6] (data from MRSA252) SACOL2215 (rpsM) 30S ribosomal protein S13 [6] (data from MRSA252) SACOL1254 (rpsP) 30S ribosomal protein S16 [6] (data from MRSA252) SACOL2230 (rpsQ) 30S ribosomal protein S17 [6] (data from MRSA252) SACOL0439 (rpsR) 30S ribosomal protein S18 [6] (data from MRSA252) SACOL2235 (rpsS) 30S ribosomal protein S19 [6] (data from MRSA252) SACOL1642 (rpsT) 30S ribosomal protein S20 [6] (data from MRSA252) SACOL2056 (rsbV) anti-anti-sigma factor RsbV [6] (data from MRSA252) SACOL2055 (rsbW) serine-protein kinase RsbW [6] (data from MRSA252) SACOL0009 (serS) seryl-tRNA synthetase [6] (data from MRSA252) SACOL0118 (sodA1) superoxide dismutase [6] (data from MRSA252) SACOL1610 (sodA2) superoxide dismutase [6] (data from MRSA252) SACOL0095 (spa) immunoglobulin G binding protein A precursor [6] (data from MRSA252) SACOL0541 (spoVG) regulatory protein SpoVG [6] (data from MRSA252) SACOL1449 (sucA) 2-oxoglutarate dehydrogenase E1 component [6] (data from MRSA252) SACOL1448 (sucB) dihydrolipoamide succinyltransferase [6] (data from MRSA252) SACOL1262 (sucC) succinyl-CoA synthetase subunit beta [6] (data from MRSA252) SACOL1263 (sucD) succinyl-CoA synthetase subunit alpha [6] (data from MRSA252) SACOL1831 (tal) translaldolase [6] (data from MRSA252) SACOL0626 (thiD1) phosphomethylpyrimidine kinase [6] (data from MRSA252) SACOL1729 (thrS) threonyl-tRNA synthetase [6] (data from MRSA252) SACOL1722 (tig) trigger factor [6] (data from MRSA252) SACOL1377 (tkt) transketolase [6] (data from MRSA252) SACOL0840 (tpiA) triosephosphate isomerase [6] (data from MRSA252) SACOL1762 (tpx) thiol peroxidase [6] (data from MRSA252) SACOL1155 (trxA) thioredoxin [6] (data from MRSA252) SACOL0829 (trxB) thioredoxin-disulfide reductase [6] (data from MRSA252) SACOL1276 (tsf) elongation factor Ts [6] (data from MRSA252) SACOL0594 (tuf) elongation factor Tu [6] (data from MRSA252) SACOL2104 (upp) uracil phosphoribosyltransferase [6] (data from MRSA252) SACOL1710 (valS) valyl-tRNA synthetase [6] (data from MRSA252) SACOL1549 (zwf) glucose-6-phosphate 1-dehydrogenase [6] (data from MRSA252) SACOL0220 flavohemoprotein [6] (data from MRSA252) SACOL0426 acetyl-CoA acetyltransferase [6] (data from MRSA252) SACOL0455 hypothetical protein [6] (data from MRSA252) SACOL0457 hypothetical protein [6] (data from MRSA252) SACOL0506 ABC transporter substrate-binding protein [6] (data from MRSA252) SACOL0521 hypothetical protein [6] (data from MRSA252) SACOL0552 hypothetical protein [6] (data from MRSA252) SACOL0564 pyridoxal biosynthesis lyase PdxS [6] (data from MRSA252) SACOL0596 2-amino-3-ketobutyrate coenzyme A ligase [6] (data from MRSA252) SACOL0599 hypothetical protein [6] (data from MRSA252) SACOL0617 hexulose-6-phosphate synthase [6] (data from MRSA252) SACOL0633 heme peroxidase [6] (data from MRSA252) SACOL0688 ABC transporter substrate-binding protein [6] (data from MRSA252) SACOL0708 DAK2 domain-containing protein [6] (data from MRSA252) SACOL0721 hypothetical protein [6] (data from MRSA252) SACOL0815 ribosomal subunit interface protein [6] (data from MRSA252) SACOL0875 thioredoxin [6] (data from MRSA252) SACOL0876 hypothetical protein [6] (data from MRSA252) SACOL0912 hypothetical protein [6] (data from MRSA252) SACOL0914 FeS assembly ATPase SufC [6] (data from MRSA252) SACOL0931 HAD superfamily hydrolase [6] (data from MRSA252) SACOL0944 NADH dehydrogenase [6] (data from MRSA252) SACOL0957 cyclophilin type peptidyl-prolyl cis-trans isomerase [6] (data from MRSA252) SACOL0973 fumarylacetoacetate hydrolase [6] (data from MRSA252) SACOL0975 coenzyme A disulfide reductase [6] (data from MRSA252) SACOL1020 hypothetical protein [6] (data from MRSA252) SACOL1099 hypothetical protein [6] (data from MRSA252) SACOL1189 hypothetical protein [6] (data from MRSA252) SACOL1240 DAK2 domain-containing protein [6] (data from MRSA252) SACOL1427 ABC transporter ATP-binding protein [6] (data from MRSA252) SACOL1437 CSD family cold shock protein [6] (data from MRSA252) SACOL1447 hypothetical protein [6] (data from MRSA252) SACOL1454 [6] (data from MRSA252) SACOL1457 PTS system, IIA component [6] (data from MRSA252) SACOL1588 proline dipeptidase [6] (data from MRSA252) SACOL1593 glycine dehydrogenase subunit 2 [6] (data from MRSA252) SACOL1594 glycine dehydrogenase subunit 1 [6] (data from MRSA252) SACOL1630 hypothetical protein [6] (data from MRSA252) SACOL1670 hypothetical protein [6] (data from MRSA252) SACOL1724 MutT/nudix family protein [6] (data from MRSA252) SACOL1749 NADP-dependent malic enzyme [6] (data from MRSA252) SACOL1753 universal stress protein [6] (data from MRSA252) SACOL1759 universal stress protein [6] (data from MRSA252) SACOL1767 septation ring formation regulator EzrA [6] (data from MRSA252) SACOL1787 bifunctional 3-deoxy-7-phosphoheptulonate synthase/chorismate mutase [6] (data from MRSA252) SACOL1789 hypothetical protein [6] (data from MRSA252) SACOL1792 hypothetical protein [6] (data from MRSA252) SACOL1794 thioredoxin [6] (data from MRSA252) SACOL1801 dipeptidase PepV [6] (data from MRSA252) SACOL1802 hypothetical protein [6] (data from MRSA252) SACOL1807 rhodanese-like domain-containing protein [6] (data from MRSA252) SACOL1891 RNAIII-activating protein TRAP [6] (data from MRSA252) SACOL1894 HIT family protein [6] (data from MRSA252) SACOL1912 hypothetical protein [6] (data from MRSA252) SACOL1952 ferritins family protein [6] (data from MRSA252) SACOL1985 hypothetical protein [6] (data from MRSA252) SACOL1992 hypothetical protein [6] (data from MRSA252) SACOL2000 hypothetical protein [6] (data from MRSA252) SACOL2114 aldehyde dehydrogenase [6] (data from MRSA252) SACOL2131 Dps family protein [6] (data from MRSA252) SACOL2163 hypothetical protein [6] (data from MRSA252) SACOL2173 alkaline shock protein 23 [6] (data from MRSA252) SACOL2288 hypothetical protein [6] (data from MRSA252) SACOL2293 NAD/NADP octopine/nopaline dehydrogenase [6] (data from MRSA252) SACOL2296 glycerate dehydrogenase [6] (data from MRSA252) SACOL2412 amino acid ABC transporter amino acid-binding protein [6] (data from MRSA252) SACOL2535 D-lactate dehydrogenase [6] (data from MRSA252) SACOL2553 pyruvate oxidase [6] (data from MRSA252) SACOL2561 hydroxymethylglutaryl-CoA synthase [6] (data from MRSA252) SACOL2569 1-pyrroline-5-carboxylate dehydrogenase [6] (data from MRSA252)
⊟Expression & Regulation[edit | edit source]
⊟Operon[edit | edit source]
- MicrobesOnline: tsf > pyrH > frr
⊟Regulation[edit | edit source]
- regulator:
⊟Transcription pattern[edit | edit source]
- S.aureus Expression Data Browser: data available for NCTC8325
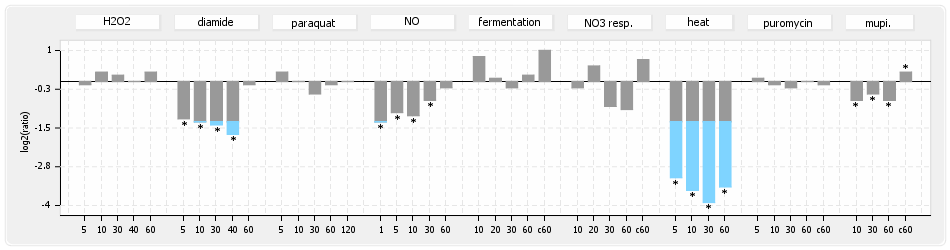
⊟Protein synthesis (provided by Aureolib)[edit | edit source]
⊟Protein stability[edit | edit source]
- half-life: no data available
⊟Biological Material[edit | edit source]
⊟Mutants[edit | edit source]
⊟Expression vector[edit | edit source]
⊟lacZ fusion[edit | edit source]
⊟GFP fusion[edit | edit source]
⊟two-hybrid system[edit | edit source]
⊟FLAG-tag construct[edit | edit source]
⊟Antibody[edit | edit source]
⊟Other Information[edit | edit source]
You can add further information about the gene and protein here. [edit]
⊟Literature[edit | edit source]
⊟References[edit | edit source]
- ↑ Dörte Becher, Kristina Hempel, Susanne Sievers, Daniela Zühlke, Jan Pané-Farré, Andreas Otto, Stephan Fuchs, Dirk Albrecht, Jörg Bernhardt, Susanne Engelmann, Uwe Völker, Jan Maarten van Dijl, Michael Hecker
A proteomic view of an important human pathogen--towards the quantification of the entire Staphylococcus aureus proteome.
PLoS One: 2009, 4(12);e8176
[PubMed:19997597] [WorldCat.org] [DOI] (I e) - ↑ Kristina Hempel, Jan Pané-Farré, Andreas Otto, Susanne Sievers, Michael Hecker, Dörte Becher
Quantitative cell surface proteome profiling for SigB-dependent protein expression in the human pathogen Staphylococcus aureus via biotinylation approach.
J Proteome Res: 2010, 9(3);1579-90
[PubMed:20108986] [WorldCat.org] [DOI] (I p) - ↑ Kristina Hempel, Florian-Alexander Herbst, Martin Moche, Michael Hecker, Dörte Becher
Quantitative proteomic view on secreted, cell surface-associated, and cytoplasmic proteins of the methicillin-resistant human pathogen Staphylococcus aureus under iron-limited conditions.
J Proteome Res: 2011, 10(4);1657-66
[PubMed:21323324] [WorldCat.org] [DOI] (I p) - ↑ Andreas Otto, Jan Maarten van Dijl, Michael Hecker, Dörte Becher
The Staphylococcus aureus proteome.
Int J Med Microbiol: 2014, 304(2);110-20
[PubMed:24439828] [WorldCat.org] [DOI] (I p) - ↑ Daniela Zühlke, Kirsten Dörries, Jörg Bernhardt, Sandra Maaß, Jan Muntel, Volkmar Liebscher, Jan Pané-Farré, Katharina Riedel, Michael Lalk, Uwe Völker, Susanne Engelmann, Dörte Becher, Stephan Fuchs, Michael Hecker
Costs of life - Dynamics of the protein inventory of Staphylococcus aureus during anaerobiosis.
Sci Rep: 2016, 6;28172
[PubMed:27344979] [WorldCat.org] [DOI] (I e) - ↑ 6.000 6.001 6.002 6.003 6.004 6.005 6.006 6.007 6.008 6.009 6.010 6.011 6.012 6.013 6.014 6.015 6.016 6.017 6.018 6.019 6.020 6.021 6.022 6.023 6.024 6.025 6.026 6.027 6.028 6.029 6.030 6.031 6.032 6.033 6.034 6.035 6.036 6.037 6.038 6.039 6.040 6.041 6.042 6.043 6.044 6.045 6.046 6.047 6.048 6.049 6.050 6.051 6.052 6.053 6.054 6.055 6.056 6.057 6.058 6.059 6.060 6.061 6.062 6.063 6.064 6.065 6.066 6.067 6.068 6.069 6.070 6.071 6.072 6.073 6.074 6.075 6.076 6.077 6.078 6.079 6.080 6.081 6.082 6.083 6.084 6.085 6.086 6.087 6.088 6.089 6.090 6.091 6.092 6.093 6.094 6.095 6.096 6.097 6.098 6.099 6.100 6.101 6.102 6.103 6.104 6.105 6.106 6.107 6.108 6.109 6.110 6.111 6.112 6.113 6.114 6.115 6.116 6.117 6.118 6.119 6.120 6.121 6.122 6.123 6.124 6.125 6.126 6.127 6.128 6.129 6.130 6.131 6.132 6.133 6.134 6.135 6.136 6.137 6.138 6.139 6.140 6.141 6.142 6.143 6.144 6.145 6.146 6.147 6.148 6.149 6.150 6.151 6.152 6.153 6.154 6.155 6.156 6.157 6.158 6.159 6.160 6.161 6.162 6.163 6.164 6.165 6.166 6.167 6.168 6.169 6.170 6.171 6.172 6.173 6.174 6.175 6.176 6.177 6.178 6.179 6.180 6.181 6.182 6.183 6.184 6.185 6.186 6.187 6.188 6.189 6.190 6.191 6.192 6.193 6.194 6.195 6.196 6.197 6.198 6.199 6.200 6.201 6.202 6.203 6.204 6.205 6.206 6.207 6.208 6.209 6.210 6.211 6.212 6.213 6.214 6.215 6.216 6.217 6.218 6.219 6.220 6.221 6.222 6.223 6.224 6.225 6.226 6.227 6.228 6.229 6.230 6.231 6.232 6.233 6.234 6.235 6.236 6.237 6.238 6.239 6.240 6.241 6.242 6.243 6.244 6.245 6.246 6.247 6.248 6.249 6.250 6.251 6.252 6.253 6.254 6.255 6.256 6.257 6.258 6.259 6.260 Artem Cherkasov, Michael Hsing, Roya Zoraghi, Leonard J Foster, Raymond H See, Nikolay Stoynov, Jihong Jiang, Sukhbir Kaur, Tian Lian, Linda Jackson, Huansheng Gong, Rick Swayze, Emily Amandoron, Farhad Hormozdiari, Phuong Dao, Cenk Sahinalp, Osvaldo Santos-Filho, Peter Axerio-Cilies, Kendall Byler, William R McMaster, Robert C Brunham, B Brett Finlay, Neil E Reiner
Mapping the protein interaction network in methicillin-resistant Staphylococcus aureus.
J Proteome Res: 2011, 10(3);1139-50
[PubMed:21166474] [WorldCat.org] [DOI] (I p)